Plot calibration results
Usage
plot_calib(
calib,
na_value,
fit_col = "fit",
nrow = 2,
base_size = 8,
return_pars = FALSE,
log_y = TRUE
)Arguments
- calib
dataframe; output from
read_calib- na_value
A numeric value which corresponds to the NA value used in the calibration.
- fit_col
character; name of column containing fit values. Default is
"fit".- nrow
integer; number of rows in plot
- base_size
numeric; base size for theme
- return_pars
logical; return parameter values
- log_y
logical; use log scale on y-axis. Default is
TRUE.
Examples
aeme_file <- system.file("extdata/aeme.rds", package = "AEME")
aeme <- readRDS(aeme_file)
model_controls <- AEME::get_model_controls()
model <- c("glm_aed", "gotm_wet")
path <- "aeme"
aeme <- AEME::build_aeme(aeme = aeme, model = model, path = path,
model_controls = model_controls, ext_elev = 5) |>
AEME::run_aeme()
#> ℹ Using observed water level
#> ℹ No missing values in observed water level. Using observed water level
#> ℹ Correcting water balance using estimated outflows (method = 2).
#> ℹ Calculating lake level using lake depth and a sinisoidal function.
#> ℹ Building GLM-AED for lake wainamu
#> ℹ Building GOTM-WET model for lake wainamu
#> ✔ GOTM YAML validation completed - no issues detected.
#> ✔ GLM nml validation completed - no issues detected.
#> ℹ Running models... (Have you tried parallelizing?) [2026-03-04 03:02:43]
#> → GLM-AED running... [2026-03-04 03:02:43]
#> ✔ GLM-AED run successful! [2026-03-04 03:02:43]
#> → GOTM-WET running... [2026-03-04 03:02:43]
#> ✔ GOTM-WET run successful! [2026-03-04 03:02:44]
#> ✔ Model run complete! [2026-03-04 03:02:44]
#> ! The following variables are not available in model gotm_wet: RAD_extc
#> ! The following variables are not available in model gotm_wet: RAD_extc
data("aeme_parameters", package = "AEME")
param <- aeme_parameters
# Function to calculate fitness
nse <- function(df) {
# Calculate Nash-Sutcliffe Efficiency
nse <- 1 - (sum((df$obs - df$model)^2) / sum((df$obs - mean(df$obs))^2))
-1 * nse
}
FUN_list <- list(HYD_temp = nse, LKE_lvlwtr = nse)
ctrl <- create_control(method = "calib", NP = 10, itermax = 20, ncore = 2,
parallel = TRUE, file_type = "db",
file_name = "results.db")
vars_sim <- c("HYD_temp", "LKE_lvlwtr")
weights <- c("HYD_temp" = 1, "LKE_lvlwtr" = 1)
# Calibrate AEME model
sim_id <- calib_aeme(aeme = aeme, model = model, path = path,
param = param, FUN_list = FUN_list, ctrl = ctrl,
vars_sim = vars_sim, weights = weights)
#> ℹ Variables not found: `LKE_lvlwtr`.
#> Adding them to model_controls.
#> ℹ Extracting indices for "glm_aed" modelled variables [2026-03-04 03:02:45]
#> ✔ Indices extracted for "glm_aed" modelled variables [2026-03-04 03:02:46]
#> ℹ Using 2 cores for parallel calibration for "glm_aed".
#> → Starting generation 1/2, 10 members. [2026-03-04 03:02:47]
#> Parameter summary for generation 1:
#> light/Kw MET_wndspd MET_radswd mixing/coef_mix_conv
#> mean 2.782 1.0080 0.9998 0.14880
#> median 2.818 1.0200 1.0020 0.14970
#> sd 1.668 0.1798 0.1893 0.03138
#> mixing/coef_wind_stir mixing/coef_mix_shear mixing/coef_mix_turb
#> mean 0.24880 0.15030 0.4591
#> median 0.24690 0.14790 0.4591
#> sd 0.03095 0.03207 0.1526
#> mixing/coef_mix_hyp outflow inflow
#> mean 0.6054 1.5120 1.4840
#> median 0.6110 1.5190 1.4980
#> sd 0.1247 0.6189 0.6108
#> ✔ Completed generation 1/2
#> for "glm_aed". [2026-03-04 03:03:01]
#> Best fit: 23.6 (sd: 3554.6) Parameters: [ 3.31, 0.784, 0.955, 0.104, 0.232,
#> 0.121, 0.538, 0.799, 0.784, and 0.798 ]
#> Writing output for generation 1 to results.db with sim ID: "45819_glmaed_C_002"
#> [2026-03-04 03:03:01]
#> ℹ Survival rate: 0.8
#> → Starting generation 2/2, 10 members. [2026-03-04 03:03:01]
#> Parameter summary for generation 2:
#> light/Kw MET_wndspd MET_radswd mixing/coef_mix_conv
#> mean 2.5380 0.9416 1.1400 0.1600
#> median 2.4260 0.9540 1.1460 0.1613
#> sd 0.6634 0.1103 0.1008 0.0305
#> mixing/coef_wind_stir mixing/coef_mix_shear mixing/coef_mix_turb
#> mean 0.21270 0.17180 0.54070
#> median 0.20770 0.17290 0.56330
#> sd 0.01511 0.02633 0.08244
#> mixing/coef_mix_hyp outflow inflow
#> mean 0.69810 1.526 1.3660
#> median 0.70170 1.554 1.2890
#> sd 0.06497 0.433 0.3933
#> Writing output for generation 2 to results.db with sim ID: "45819_glmaed_C_002"
#> [2026-03-04 03:03:07]
#> ✔ Completed generation 2/2
#> for "glm_aed". [2026-03-04 03:03:07]
#> Best fit: 23.6 (sd: 2212.7)
#> ℹ Survival rate: 0.9
#> ℹ Extracting indices for "gotm_wet" modelled variables [2026-03-04 03:03:07]
#> ✔ Indices extracted for "gotm_wet" modelled variables [2026-03-04 03:03:08]
#> ℹ Using 2 cores for parallel calibration for "gotm_wet".
#> → Starting generation 1/2, 10 members. [2026-03-04 03:03:09]
#> Parameter summary for generation 1:
#> turbulence/turb_param/k_min light_extinction/A/constant_value
#> mean 4.979e-06 0.52760
#> median 4.860e-06 0.52320
#> sd 3.017e-06 0.08128
#> light_extinction/g1/constant_value light_extinction/g2/constant_value
#> mean 0.58820 1.3750
#> median 0.59020 1.3230
#> sd 0.09361 0.8002
#> MET_wndspd MET_radswd outflow inflow
#> mean 0.9927 0.9937 1.5220 1.5040
#> median 0.9915 0.9995 1.5320 1.5070
#> sd 0.1844 0.1758 0.5937 0.5998
#> ✔ Completed generation 1/2
#> for "gotm_wet". [2026-03-04 03:03:28]
#> Best fit: 59.4 (sd: 11101) Parameters: [ 2.63e-06, 0.439, 0.709, 1.74, 0.954,
#> 0.739, 1.04, and 0.975 ]
#> Writing output for generation 1 to results.db with sim ID:
#> "45819_gotmwet_C_002" [2026-03-04 03:03:28]
#> ℹ Survival rate: 0.6
#> → Starting generation 2/2, 10 members. [2026-03-04 03:03:28]
#> Parameter summary for generation 2:
#> turbulence/turb_param/k_min light_extinction/A/constant_value
#> mean 4.248e-06 0.4881
#> median 3.681e-06 0.4879
#> sd 2.642e-06 0.0791
#> light_extinction/g1/constant_value light_extinction/g2/constant_value
#> mean 0.65040 1.3980
#> median 0.66210 1.5400
#> sd 0.09217 0.5722
#> MET_wndspd MET_radswd outflow inflow
#> mean 0.92330 0.8062 1.1730 1.2310
#> median 0.92010 0.7762 1.1890 1.2180
#> sd 0.04755 0.1147 0.2349 0.3527
#> Writing output for generation 2 to results.db with sim ID:
#> "45819_gotmwet_C_002" [2026-03-04 03:03:39]
#> ✔ Completed generation 2/2
#> for "gotm_wet". [2026-03-04 03:03:39]
#> Best fit: 39 (sd: 3732)
#> ℹ Survival rate: 1
# Read calibration output
calib <- read_calib(sim_id = sim_id, ctrl = ctrl)
plist <- plot_calib(calib = calib)
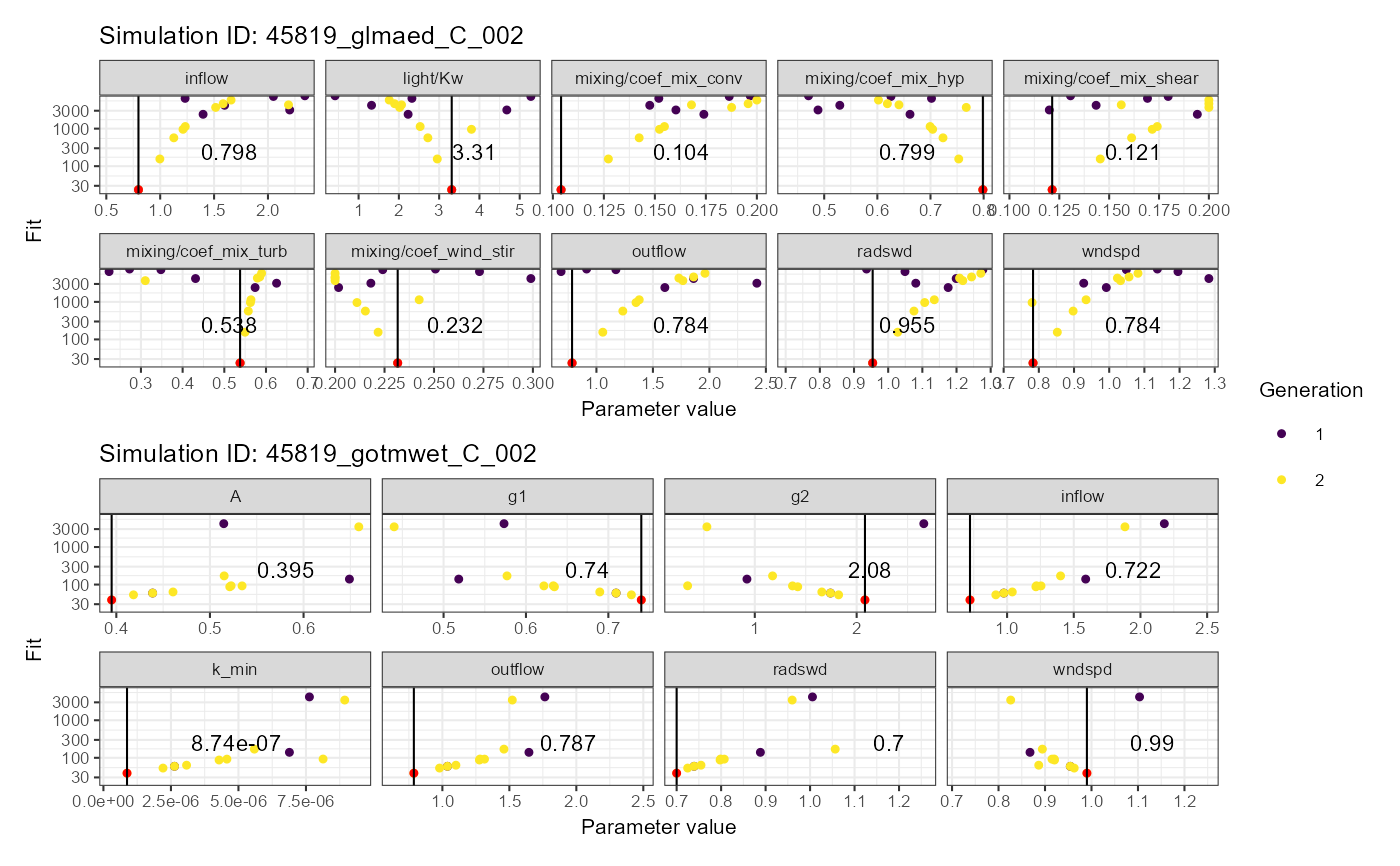
# Dotty plot
plist$dotty
#> Warning: Removed 30 rows containing missing values or values outside the scale range
#> (`geom_point()`).
#> Warning: Removed 32 rows containing missing values or values outside the scale range
#> (`geom_point()`).
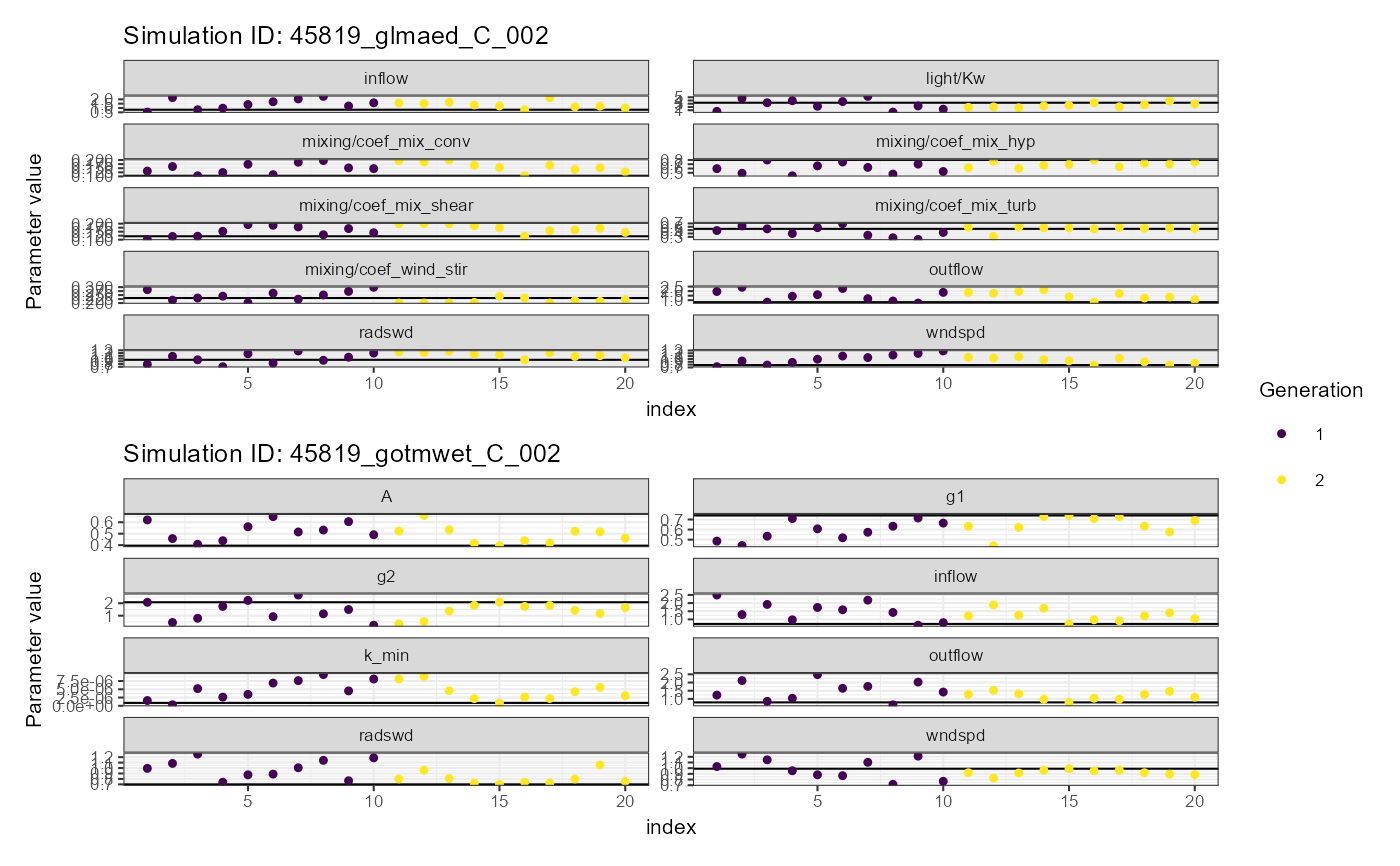
 # Convergence plot
plist$convergence
# Convergence plot
plist$convergence
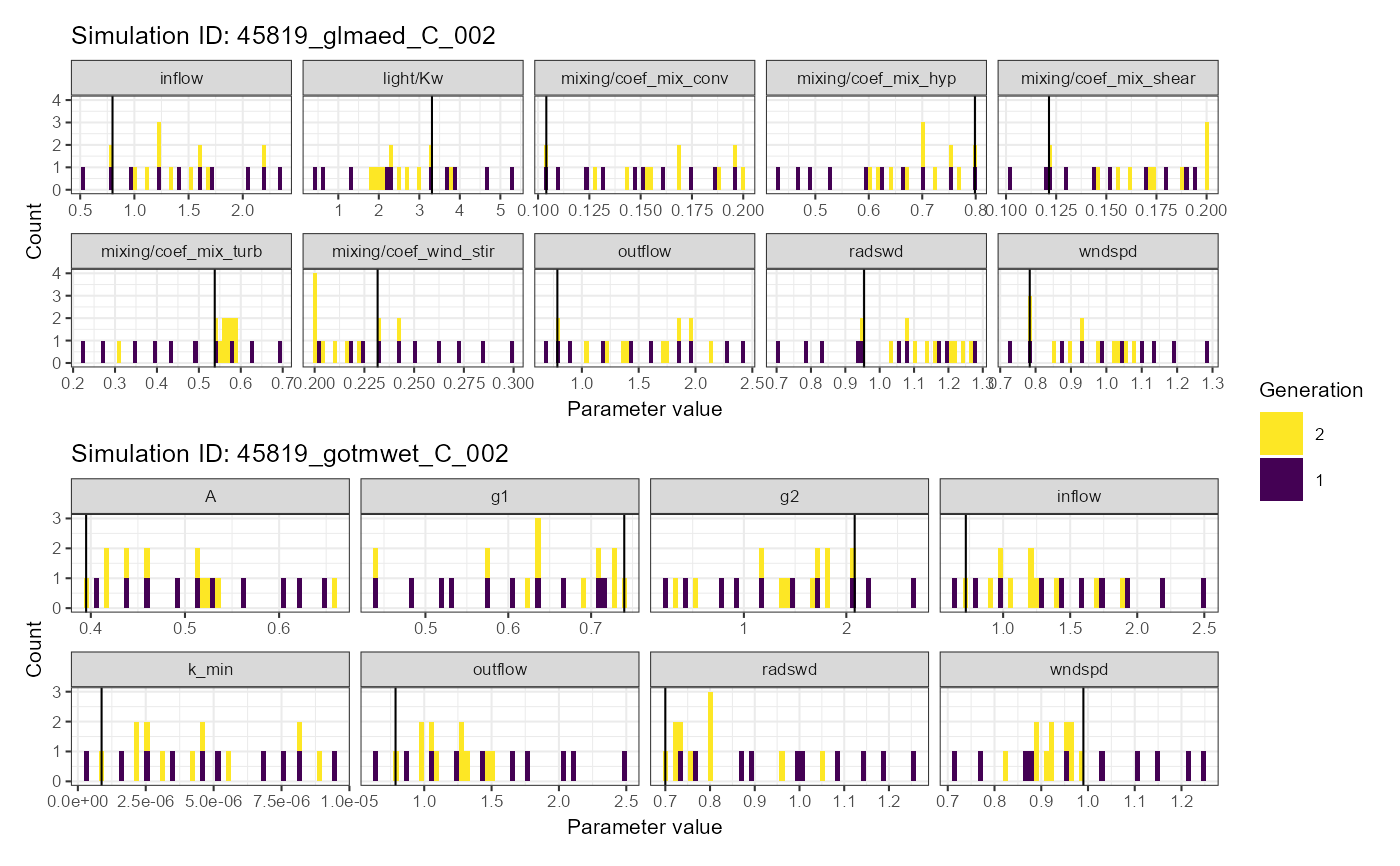
 # Histogram plot
plist$histogram
# Histogram plot
plist$histogram