Load data
The raw data was downloaded from the Bay of Plenty hydroTel SFTP server. The data were then collated into a CSV file which can be downloaded from GitHub or within R using the piggyback package. The data are stored in the data-raw folder.
<- TRUE if (download) {dir.create ("data-raw" , showWarnings = FALSE )# Hydrotel data :: pb_download (file = "rotoiti_data_raw.zip" ,dest = "." ,repo = "limnotrack/f_rotoiti" ,tag = "v0.0.1" , overwrite = TRUE # Unzip the file to tempdir <- tempdir ()unzip ("rotoiti_data_raw.zip" , exdir = tmp_dir)# Move files into data-raw file.copy (list.files (file.path (tmp_dir, "rotoiti_data_raw" ), full.names = TRUE ),"data-raw" , overwrite = TRUE )# Delete zip file unlink ("rotoiti_data_raw.zip" , force = TRUE )# Read in raw data <- read_csv ("data-raw/rotoiti_sftp_compiled_raw.csv" ) |> mutate (datetime = as_datetime (Date)) |> select (- Date) |> arrange (datetime)
Rows: 467194 Columns: 24
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
dbl (23): battery_voltage, chlorophyll_1m, compass_heading, dir, do_conc_18...
dttm (1): Date
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.
Site location
Lake Rotoiti is located in the Bay of Plenty region of New Zealand. The monitoring site is located near the centre of the lake around its deepest point.
Compile raw hydroTel data from BoP
Check no variables are time-aggregated
Standardise to 15-minute data (i.e. modify any periods where data were logged every 5-minutes.
Circular average for wind direction, sum for rainfall, mean for everything else.
<- dat |> select (datetime, contains ("dir" )) |> standardise (15 , FUN = avg_circ)<- c ("datetime" , "do_conc_1m" , "do_conc_18m" ,"do_sat_1m" , "do_sat_18m" ,"ph_decommissioned" , "ph_raw" ,"temp_0m" , "temp_2m" , "temp_4m" , "temp_6m" , "temp_9m" ,"temp_12m" , "temp_15m" , "temp_18m" , "temp_21m" ,"chlorophyll_1m" , "wind_speed_m-s" , "wind_speed" )<- dat |> select (contains (mean_vars)) |> standardise (15 , FUN = mean, na.rm = TRUE )<- data.frame (datetime = seq.POSIXt (round_date (min (dat$ datetime), "15 mins" ),round_date (max (dat$ datetime), "15 mins" ),by = 15 * 60 ))<- list (date_df, means, wind_dir) |> # left_join(means, by = "datetime") |> :: reduce (function (x, y) {left_join (x, y, by = "datetime" )}) |> arrange (datetime)
Map column names
Map column names to standard names used in the database.
<- c ("f_chl_d100" = "chlorophyll_1m" ,"c_do_d100" = "do_conc_1m" ,"c_do_d1800" = "do_conc_18m" ,"c_do_sat_d100" = "do_sat_1m" ,"c_do_sat_d1800" = "do_sat_18m" ,"t_wtr_d50" = "temp_0m" ,"t_wtr_d200" = "temp_2m" ,"t_wtr_d400" = "temp_4m" ,"t_wtr_d600" = "temp_6m" ,"t_wtr_d900" = "temp_9m" ,"t_wtr_d1200" = "temp_12m" ,"t_wtr_d1500" = "temp_15m" ,"t_wtr_d1800" = "temp_18m" ,"t_wtr_d1810" = "do_temp_18m" ,"t_wtr_d2100" = "temp_21m" ,"w_spd_h150" = "wind_speed_m-s" ,"w_dir_h150" = "dir" )<- standardise_columns (dat2, col_mapping) |> select (- wind_speed)
Write level one raw data to file.
write_csv (dat3, "data-raw/rotoiti_sftp_compiled_raw_level1.csv" )
Sensor Attribution
Map each sensor onto each data point.
Convert wide dataframe to standardised long format.
<- raw |> pivot_longer (cols = - datetime, names_to = "var_ref_id" , values_to = "raw_value" ) <- raw_long |> select (var_ref_id) |> distinct () |> arrange (var_ref_id) |> # rowwise() |> mutate (var_abbr = decode_var_ref (var_ref = var_ref_id)$ var_abbr) # Conditional join raw_long with sensor_map using var_ref_id and when datetime is between date_from and date_to # raw_long_device <- sensor_map |> # # slice(1:3) |> # group_by(device_id) |> # left_join(raw_long, by = "var_ref_id") |> # filter(datetime >= date_from & datetime <= date_to) |> # ungroup() |> # arrange(device_id, datetime) |> <- raw_long |> mutate (site = "f_Rotoiti" ,device_id = case_when (%in% c ("t_wtr_d1810" , "c_do_d1800" , "c_do_sat_d1800" ) ~ "AA_device_c_do_d1800" ,%in% c ("c_do_d100" , "c_do_sat_d100" ) ~ "AA_device_c_do_d100" ,grepl ("t_wtr" , var_ref_id) ~ "AA_device_t_wtr_string" ,.default = paste0 ("AA_device_" , var_ref_id)|> left_join (vars_abbr, by = "var_ref_id" ) |> mutate (qc_value = round (raw_value, 2 ),qc_flag = "" , qc_code = case_when (is.na (raw_value) ~ "QC 100" ,TRUE ~ "QC 200" |> select (site, datetime, device_id, var_abbr, var_ref_id, raw_value, qc_value, |> distinct (datetime, var_ref_id, .keep_all = TRUE ) <- extract_device_metadata (raw_long_device)<- site_meta$ site_devices<- site_meta$ device_position<- site_meta$ device_var<- map_sensors (site_devices = site_devices,device_var = device_var,device_position = device_position)head (raw_long_device)
# A tibble: 6 × 9
site datetime device_id var_abbr var_ref_id raw_value qc_value
<chr> <dttm> <chr> <chr> <chr> <dbl> <dbl>
1 f_Rotoiti 2007-01-12 05:00:00 AA_devic… c_do c_do_d100 0.41 0.41
2 f_Rotoiti 2007-01-12 05:00:00 AA_devic… c_do c_do_d1800 NaN NaN
3 f_Rotoiti 2007-01-12 05:00:00 AA_devic… c_do_sat c_do_sat_… NaN NaN
4 f_Rotoiti 2007-01-12 05:00:00 AA_devic… c_do_sat c_do_sat_… 1.39 1.39
5 f_Rotoiti 2007-01-12 05:00:00 AA_devic… t_wtr t_wtr_d50 NaN NaN
6 f_Rotoiti 2007-01-12 05:00:00 AA_devic… t_wtr t_wtr_d200 NaN NaN
# ℹ 2 more variables: qc_flag <chr>, qc_code <chr>
Remove based on site events
# raw_long_device <- raw_long_device |> # remove_site_events(site_events = site_events, sensor_map = sensor_map) # # raw_long_device |> # filter(qc_flag == "sensor_fault") # plot_qc_data(data = raw_long_device_filt)
Quality control
We used a set of quality control codes to assess the quality of the data. The codes are from the National Environmental Monitoring Standards (NEMS) . The codes are as follows:
<- c ("Missing Record" = "QC 100" ,"No Quality or Non Verified" = "QC 200" ,"Synthetic" = "QC 300" ,"Poor Quality" = "QC 400" ,"Fair Quality" = "QC 500" ,"Good Quality" = "QC 600" )= c ("QC 100" = "#FF0000" ,"QC 200" = "#8B5A00" ,"QC 300" = "#D3D3D3" ,"QC 400" = "#FFA500" ,"QC 500" = "#00BFFF" ,"QC 600" = "#006400" <- data.frame (qc_code = qc_codes,qc_zone = names (qc_codes),qc_code_col = unname (qc_code_col_scale),stringsAsFactors = FALSE |> datatable (rownames = FALSE ,options = list (pageLength = 6 ,dom = "t" ,columnDefs = list (list (className = 'dt-center' , targets = "_all" ))|> formatStyle ('qc_code_col' ,target = 'row' ,backgroundColor = styleEqual (qc_code_col_scale, qc_code_col_scale)
Basic QC - filters
Filter extreme outliers (order of magnitude plus)
Filter repetitions/stuck values (careful genuine repeats e.g., rainfall == 0)
Remove data for any periods where buoy was clearly out of the water (see notes or visual check of data)
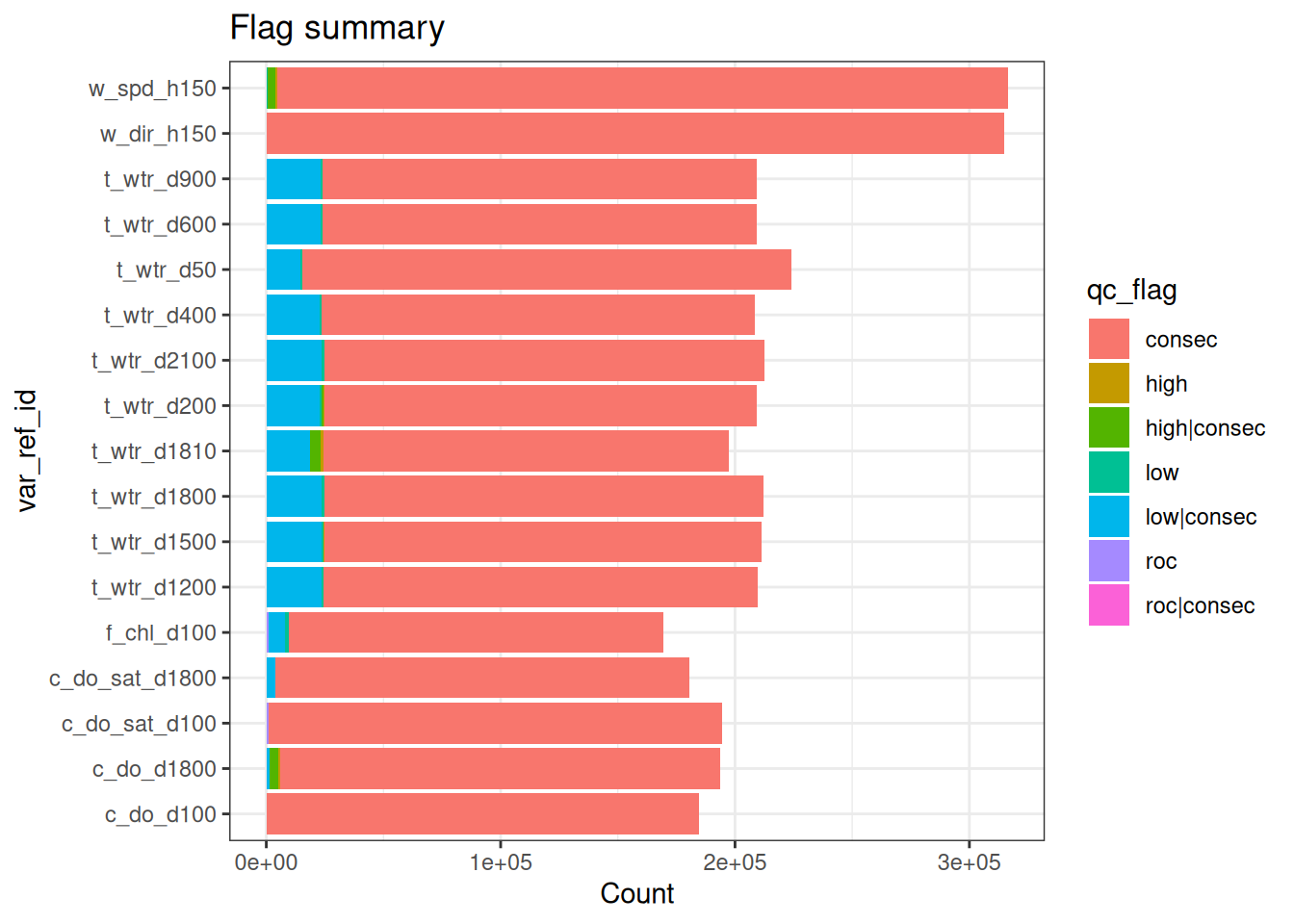
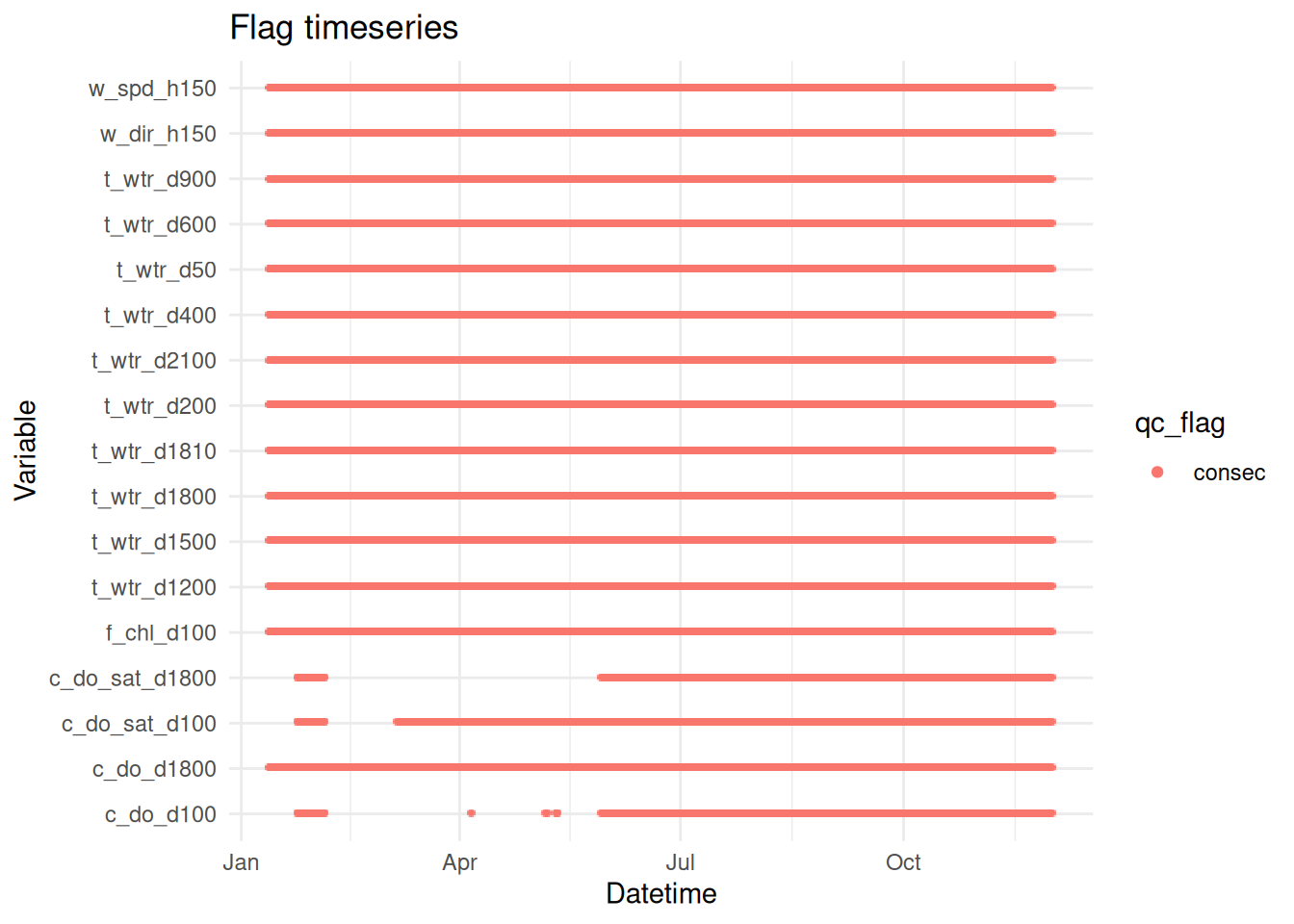
<- apply_filters (raw_long_device2, filters = qc_filters)
plot_flag_ts (raw_long_device_filt)
Fluorescence sensors
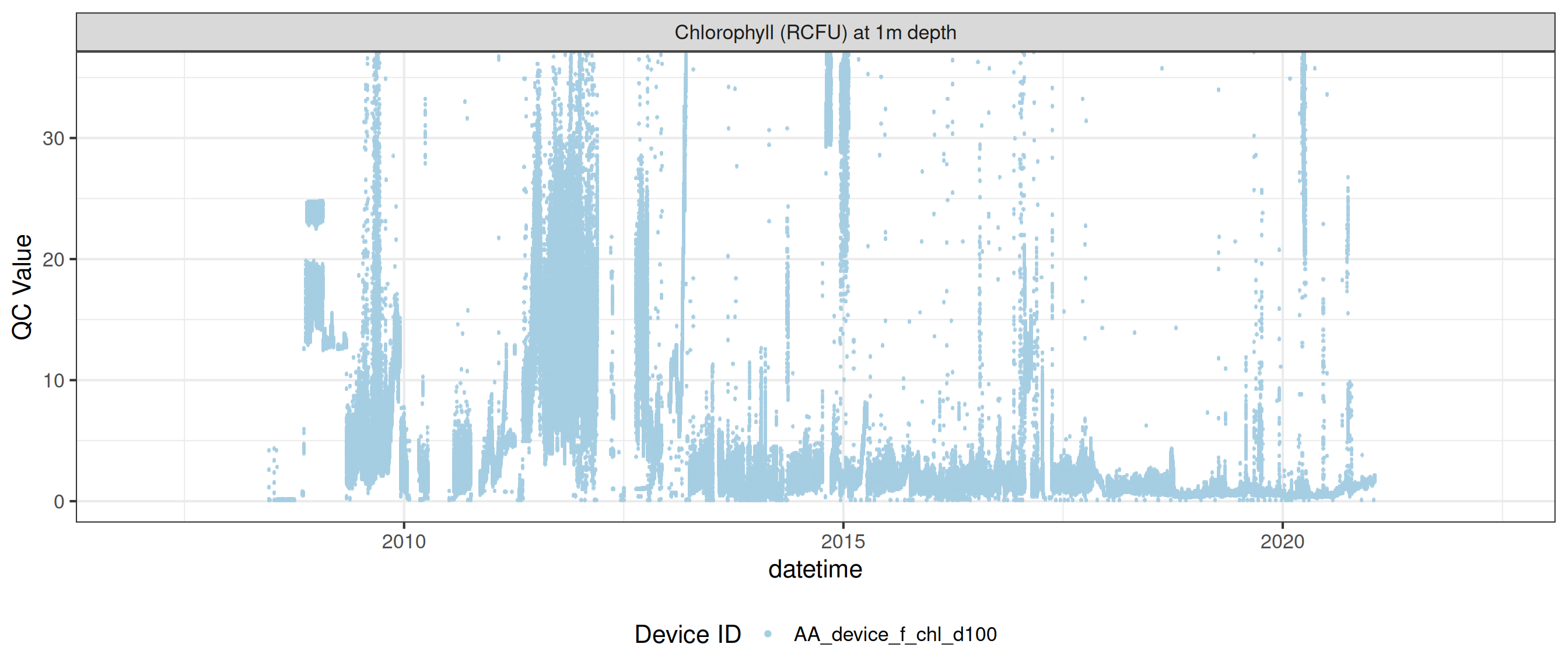
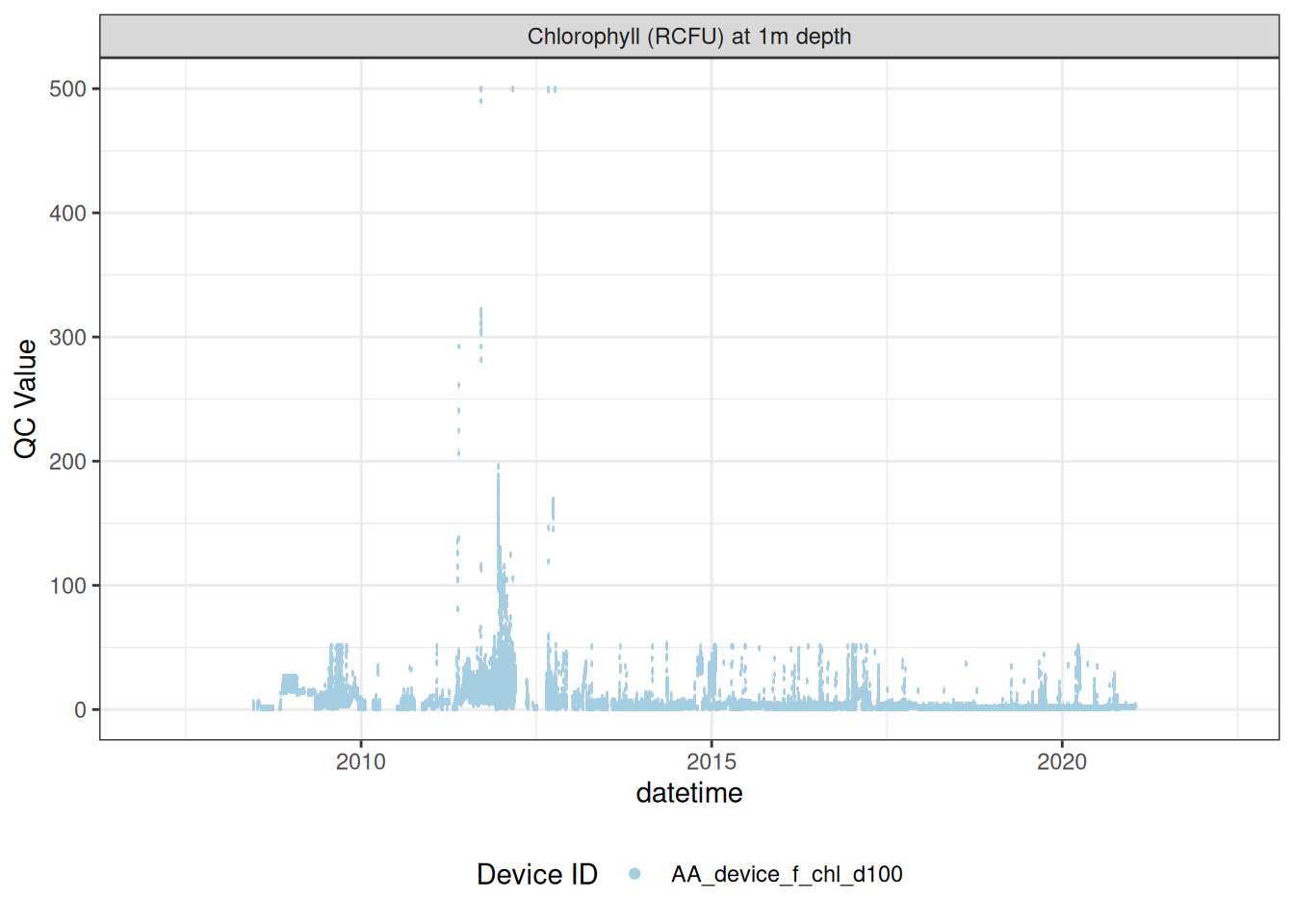
<- raw_long_device_filt |> filter (var_abbr %in% c ("f_chl" )) #|> # apply_adjustment("f_chl_d100", # date_range = c("2008-11-15 23:23:03", "2009-01-29 12:59:29"), # FUN = \(x) x / 5) |> # apply_adjustment("f_chl_d100", # date_range = c("2009-01-29 13:00:00", "2011-04-20 17:30:00"), # FUN = \(x) x / 10) |> # apply_adjustment("f_chl_d100", # date_range = c("2011-04-20 17:30:34", "2012-10-11 21:18:38"), # FUN = \(x) x / 100) |> # apply_adjustment("f_chl_d100", # date_range = c("2012-10-11 21:19:38", "2021-02-19 03:55:08"), # FUN = \(x) x / 10) |> # apply_adjustment("f_chl_d100", # FUN = \(x) ifelse(x >= 5, NA, x))# |> # viz_data() # fluor_sensors <- sensor_map |> # filter(device_id %in% fluor_df$device_id) |> # select(device_id, var_ref_id, date_from, date_to) # # sensor_refs <- sensor_reference |> # left_join(fluor_sensors, by = "device_id") |> # filter(device_id %in% fluor_sensors$device_id, # !is.na(value_actual) # # date > "2013-06-28", # # value_actual %in% c(4, 7, 10), # # units_sensor == "counts" # ) # # sensor_scales <- sensor_scaling |> # arrange(date) |> # filter(device_id %in% fluor_sensors$device_id) plot_sensor (data = fluor_df, var_ref_id = c ("f_chl_d100" ),#, "f_phyc_d100"), variable_ref = variable_ref, sensor_calibrations = sensor_calibrations, sensor_reference = sensor_reference,sensor_scaling = sensor_scaling, clip_ylim = F)
Correct fluorescence data by removing bad data and applying corrections to get the chlorophyll at it its raw 0-5V raw signal, then recorrecting to get the corrected chlorophyll value inrelative fluorescene units (RFU).
<- fluor_df |> filter (var_ref_id == "f_chl_d100" )# viz_data(data = raw_chl, site_events = site_events, variable = "var_ref_id", # value = "qc_value") # sensor_scales <- sensor_scaling |> # filter(device_id %in% raw_chl$device_id) |> # select(device_id, date, offset, multiplier, range) <- raw_chl |> mutate (var_ref_id = "f_chl_raw_v_d100" ,var_abbr = "f_chl_raw_v" ,# raw_value = NA_real_ |> apply_adjustment ("f_chl_raw_v_d100" ,date_range = c ("2008-11-15 23:23:03" , "2009-01-29 12:59:29" ),FUN = \(x) x / 5 ) |> apply_adjustment ("f_chl_raw_v_d100" ,date_range = c ("2009-05-05 12:31:33" , "2009-12-16 04:43:55" ),FUN = \(x) x / 3 ) |> apply_adjustment ("f_chl_raw_v_d100" ,date_range = c ("2009-01-29 13:00:00" , "2011-04-20 17:30:00" ),FUN = \(x) x / 10 ) |> apply_adjustment ("f_chl_raw_v_d100" ,date_range = c ("2011-04-20 17:30:34" , "2012-10-11 21:18:38" ),FUN = \(x) x / 100 ) |> apply_adjustment ("f_chl_raw_v_d100" ,date_range = c ("2012-10-11 21:19:38" , "2021-02-19 03:55:08" ),FUN = \(x) x / 10 ) |> apply_adjustment ("f_chl_raw_v_d100" ,FUN = \(x) ifelse (x >= 5 , NA , x)) |> update_qc_code_vectorized (qc_update_df = qc_update_df)
<- raw_v |> # update_qc_code_vectorized(qc_update_df = qc_update_df) |> mutate (qc_value = qc_value * 10 ,var_ref_id = "f_chl_d100" ,var_abbr = "f_chl" ,|> plot_sensor (var_ref_id = "f_chl_d100" , variable_ref = variable_ref, sensor_calibrations = sensor_calibrations, sensor_reference = sensor_reference,sensor_scaling = sensor_scaling, clip_ylim = FALSE ,colour = "qc_code" )
Load in field measurements
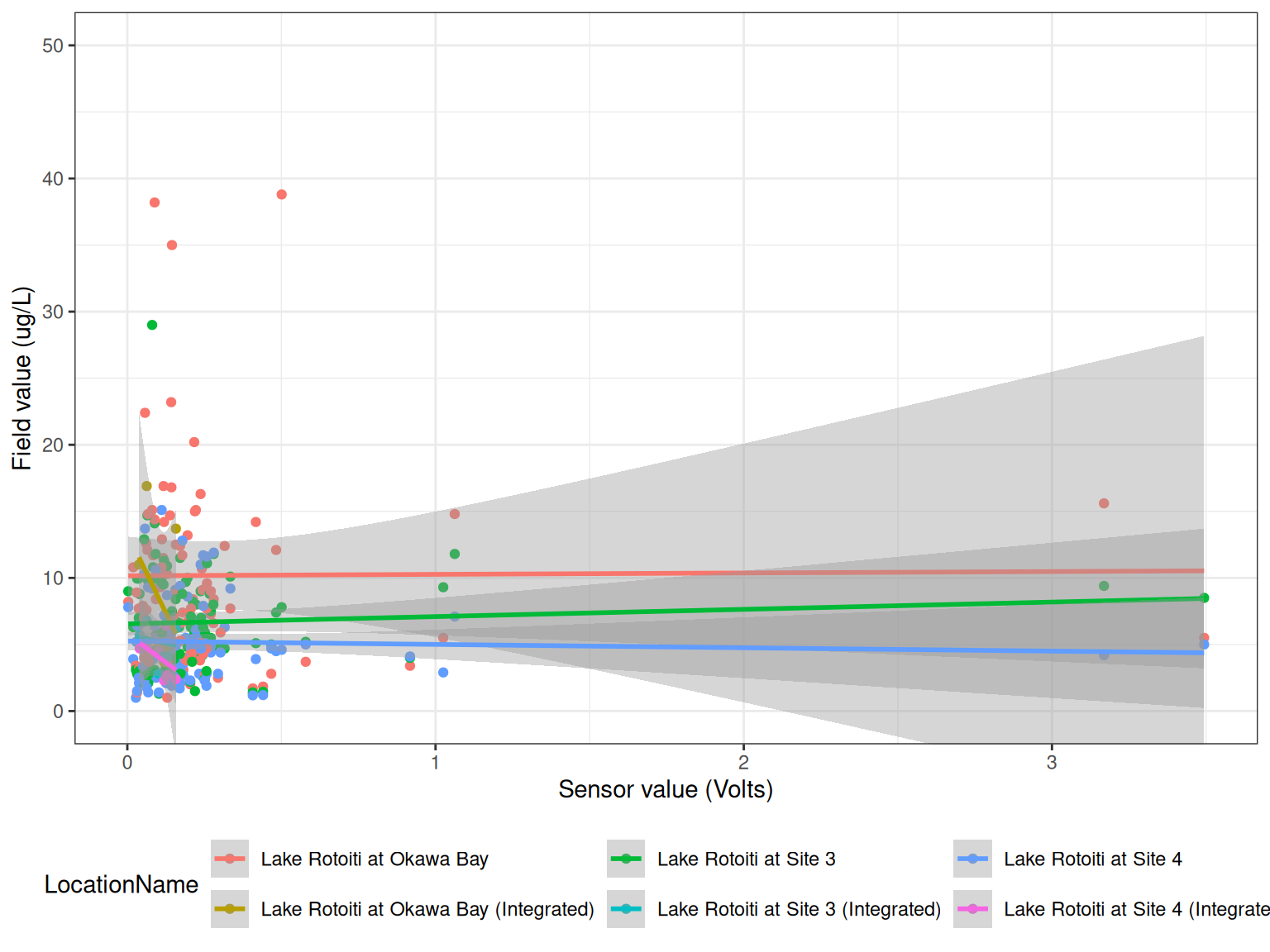
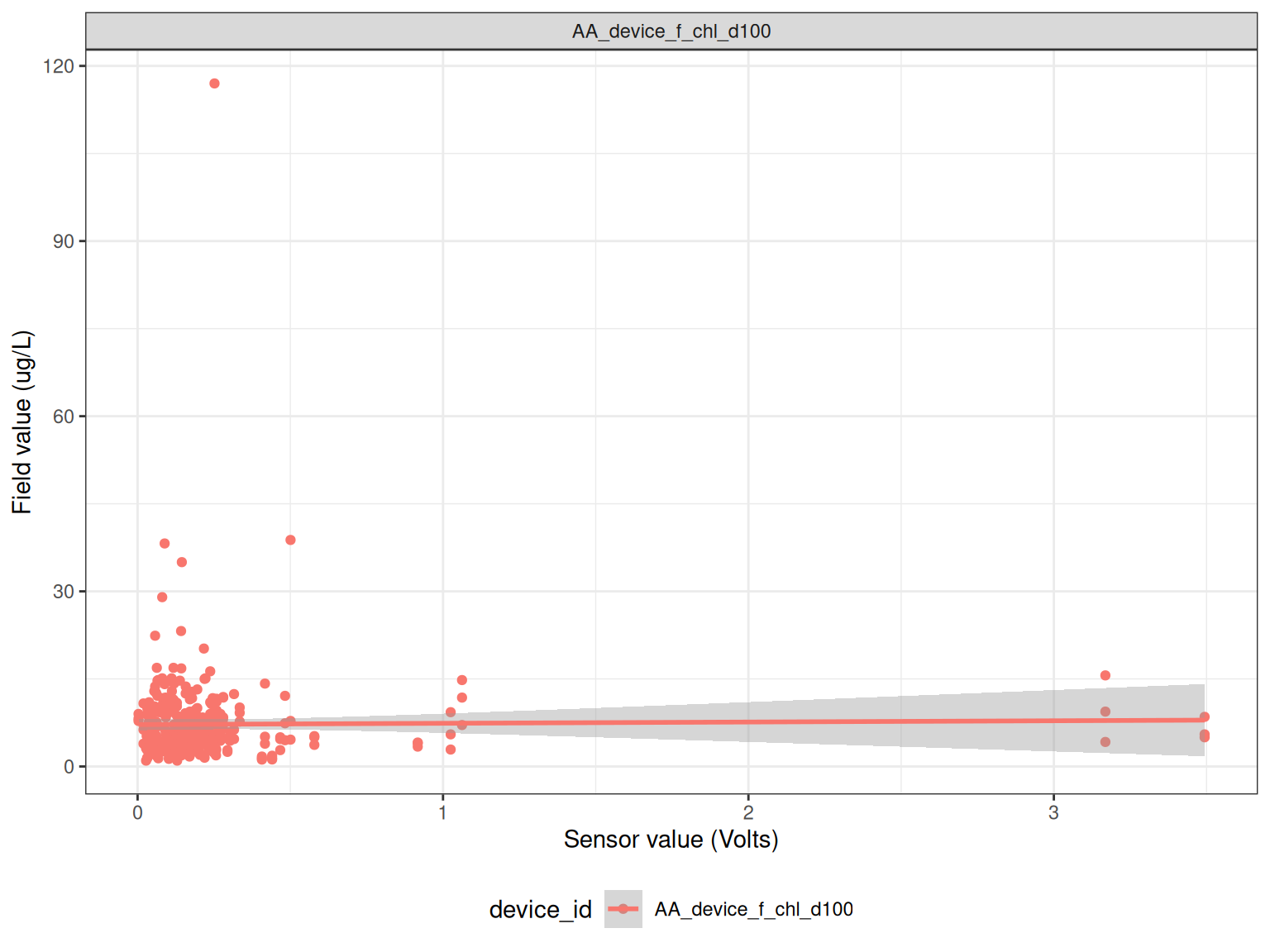
Compare field measurements to sensor data.
<- read_csv ("data-raw/rotoiti_field_data.csv" ) |> mutate (Date2 = as.Date (Date))<- field |> filter (grepl ("Chla" , Parameter)) |> select (Date2, LocationName, DepthFrom, Sample_Depth, Value) |> rename (Date = Date2)<- raw_long_device_filt |> mutate (hour = hour (datetime)) |> filter (var_abbr %in% c ("f_chl" )) |> filter (hour %in% c (0 : 6 , 21 : 23 )) |> filter (as.Date (datetime) %in% as.Date (chla$ Date))<- raw_v |> mutate (Date = as.Date (datetime)) |> filter (datetime %in% ref_times$ datetime) |> group_by (Date, device_id) |> summarise (qc_value = mean (qc_value, na.rm = TRUE ),median = median (qc_value, na.rm = TRUE ), .groups = "drop" )<- sub_sensor |> left_join (chla, by = c ("Date" = "Date" )) |> mutate (year = year (Date)) |> filter (! is.na (qc_value))ggplot () + geom_point (data = df, aes (qc_value, Value, colour = LocationName)) + geom_smooth (data = df, aes (qc_value, Value, colour = LocationName), method = "lm" ) + # facet_wrap(year~device_id, scales = "free") + labs (x = "Sensor value (Volts)" ,y = "Field value (ug/L)" ) + coord_cartesian (ylim = c (0 , 50 )) + theme_bw () + theme (legend.position = "bottom" ) ggplot () + geom_point (data = df, aes (qc_value, Value, colour = device_id)) + geom_smooth (data = df, aes (qc_value, Value, colour = device_id), method = "lm" ) + facet_wrap (~ device_id, scales = "free" ) + labs (x = "Sensor value (Volts)" ,y = "Field value (ug/L)" ) + # coord_cartesian(ylim = c(0, 50)) + theme_bw () + theme (legend.position = "bottom" )
Oxygen sensors
Remove bad data and correct for linear drift
<- raw_long_device_filt |> filter (var_ref_id %in% c ("c_do_sat_d100" )) |> drift_correction (var_ref_id = "c_do_sat_d100" , date_range = c ("2017-05-13 12:29:20" , "2020-11-09 23:18:35" ), low = c (0 , 0 , 0 ), high = c (100 , 88 , 88 )) |> update_qc_code_vectorized (qc_update_df = qc_update_df)
plot_raw_qc (data = do_1m, variable_ref = variable_ref, ylim = c (80 , 150 ))# do_1m |> # viz_data(qc_update_df = qc_update_df, site_events = site_events, # variable = "var_ref_id", value = "qc_value")
|> # plotly_data(y1 = "c_do_sat_d100", sub = 20) plot_sensor (var_ref_id = c ("c_do_sat_d100" ), variable_ref = variable_ref, sensor_calibrations = sensor_calibrations, sensor_reference = sensor_reference,sensor_scaling = sensor_scaling, clip_ylim = FALSE ,colour = "qc_code" )
<- raw_long_device_filt |> filter (var_ref_id %in% c ("c_do_sat_d1800" )) #|> # update_qc_code_vectorized(qc_update_df = qc_update_df)
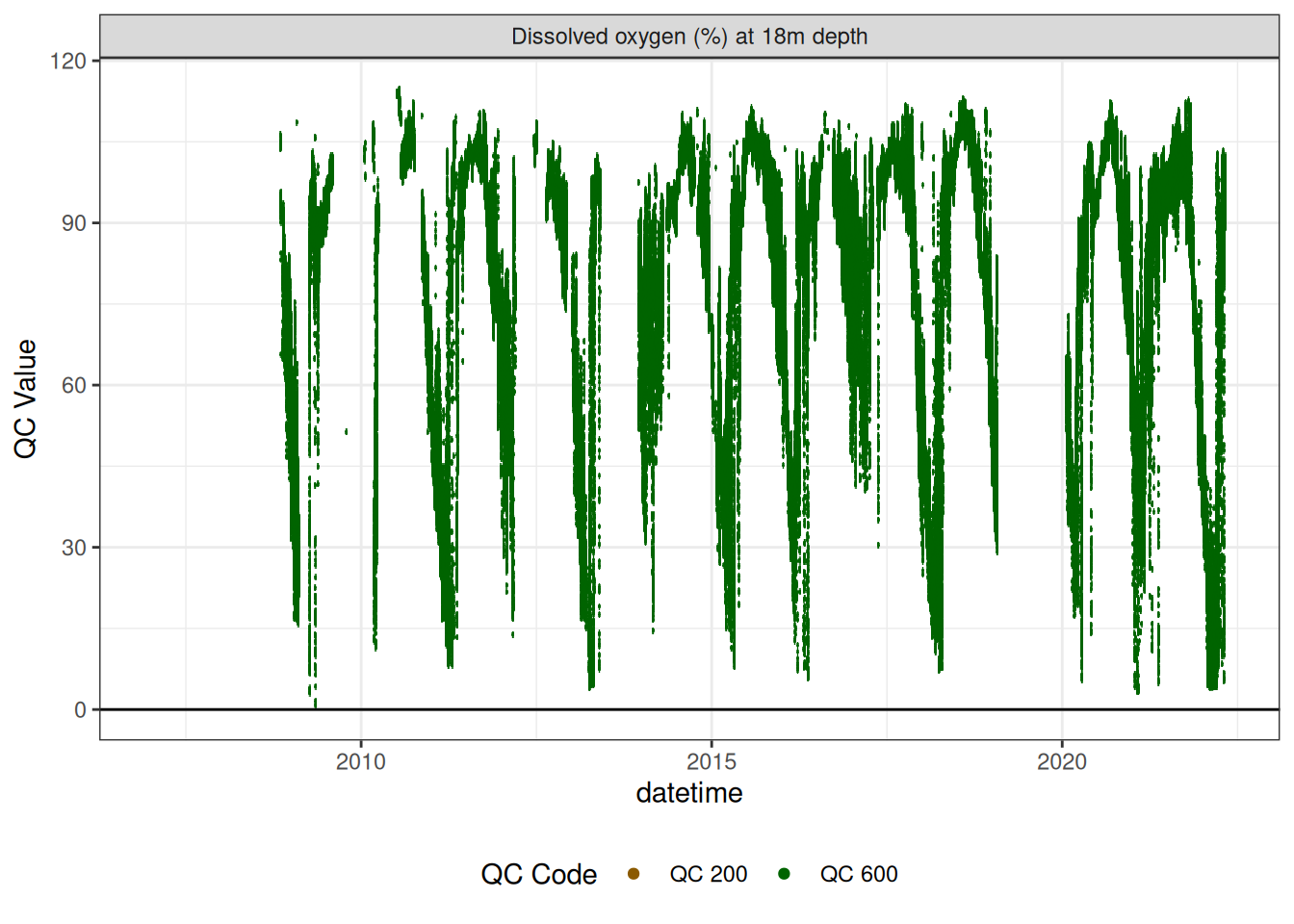
<- do_18m |> update_qc_code_vectorized (qc_update_df = qc_update_df) plot_raw_qc (data = do_18m, variable_ref = variable_ref, ylim = c (0 , 120 ))
|> # plotly_data(y1 = "c_do_sat_d1000", sub = 20) plot_sensor (var_ref_id = c ("c_do_sat_d1800" ), variable_ref = variable_ref,sensor_calibrations = sensor_calibrations, sensor_reference = sensor_reference,sensor_scaling = sensor_scaling, clip_ylim = F, colour = "qc_code" ) + geom_hline (yintercept = 0 )
Recalculate DO concentrations using temperature and DO saturation
Download the Rotoehu data quality control file from the repository and unzip it. We will use the air pressure to calculate the DO saturation concentration.
<- raw_long_device_filt |> filter (var_ref_id == "t_wtr_d50" )
<- temp_1m |> update_qc_code_vectorized (qc_update_df = qc_update_df) |> select (datetime, qc_value) |> rename (temp = qc_value)<- data_ehu |> filter (var_ref_id == "pr_baro_h150" ) |> select (datetime, qc_value) |> rename (pr_baro = qc_value)<- left_join (do_1m, rel_temp, by = "datetime" ) |> left_join (pr_baro, by = "datetime" ) |> mutate (do_sat_mgL = calc_DOsat_mg (temp, pr_baro),var_abbr = "c_do" ,var_ref_id = "c_do2_d100" ,qc_value = qc_value / 100 * do_sat_mgL|> select (datetime, var_ref_id, site, device_id, var_abbr, qc_value, qc_flag, <- raw_long_device_filt |> filter (var_ref_id == "c_do_d100" ) |> select (datetime, var_ref_id, site, device_id, var_abbr, raw_value)<- do_conc2_1m |> mutate (var_ref_id = "c_do_d100" ) |> select (datetime, var_ref_id, site, device_id, var_abbr, qc_value,<- left_join (do_conc_1m, do_conc2_1m,by = c ("datetime" , "var_ref_id" , "site" , "device_id" , "var_abbr" )) |> mutate (qc_code = case_when (! is.na (qc_value) ~ "QC 300" ,is.na (raw_value) ~ "QC 100" ,is.na (qc_value) ~ "QC 200"
<- raw_long_device_filt |> filter (var_ref_id == "t_wtr_d1810" )
<- temp_18m |> update_qc_code_vectorized (qc_update_df = qc_update_df) <- temp_18m |> select (datetime, qc_value) |> rename (temp = qc_value)<- left_join (do_18m, temp_18m, by = "datetime" ) |> left_join (pr_baro, by = "datetime" ) |> mutate (do_sat_mgL = calc_DOsat_mg (temp, pr_baro),var_abbr = "c_do" ,var_ref_id = "c_do2_d100" ,qc_value = qc_value / 100 * do_sat_mgL|> select (datetime, var_ref_id, site, device_id, var_abbr, qc_value, qc_flag, do_sat_mgL, temp)<- raw_long_device_filt |> filter (var_ref_id == "c_do_d1800" ) |> select (datetime, var_ref_id, site, device_id, var_abbr, raw_value)<- do_conc2_18m |> mutate (var_ref_id = "c_do_d1800" ) |> select (datetime, var_ref_id, site, device_id, var_abbr, qc_value,<- left_join (do_conc_18m, do_conc2_18m, by = c ("datetime" , "var_ref_id" , "site" , "device_id" , "var_abbr" )) |> mutate (qc_code = case_when (! is.na (qc_value) ~ "QC 300" ,is.na (raw_value) ~ "QC 100" ,is.na (qc_value) ~ "QC 200"
plot_raw_qc (data = do_conc3_18m, variable_ref = variable_ref, ylim = c (0 , 12 ))
<- bind_rows (do_conc3_1m, do_conc3_18m, do_1m, do_18m) # qc_update_df <- viz_data(data = do, qc_update_df = qc_update_df, # site_events = site_events, variable = "var_ref_id", # value = "qc_value")
Water temperature
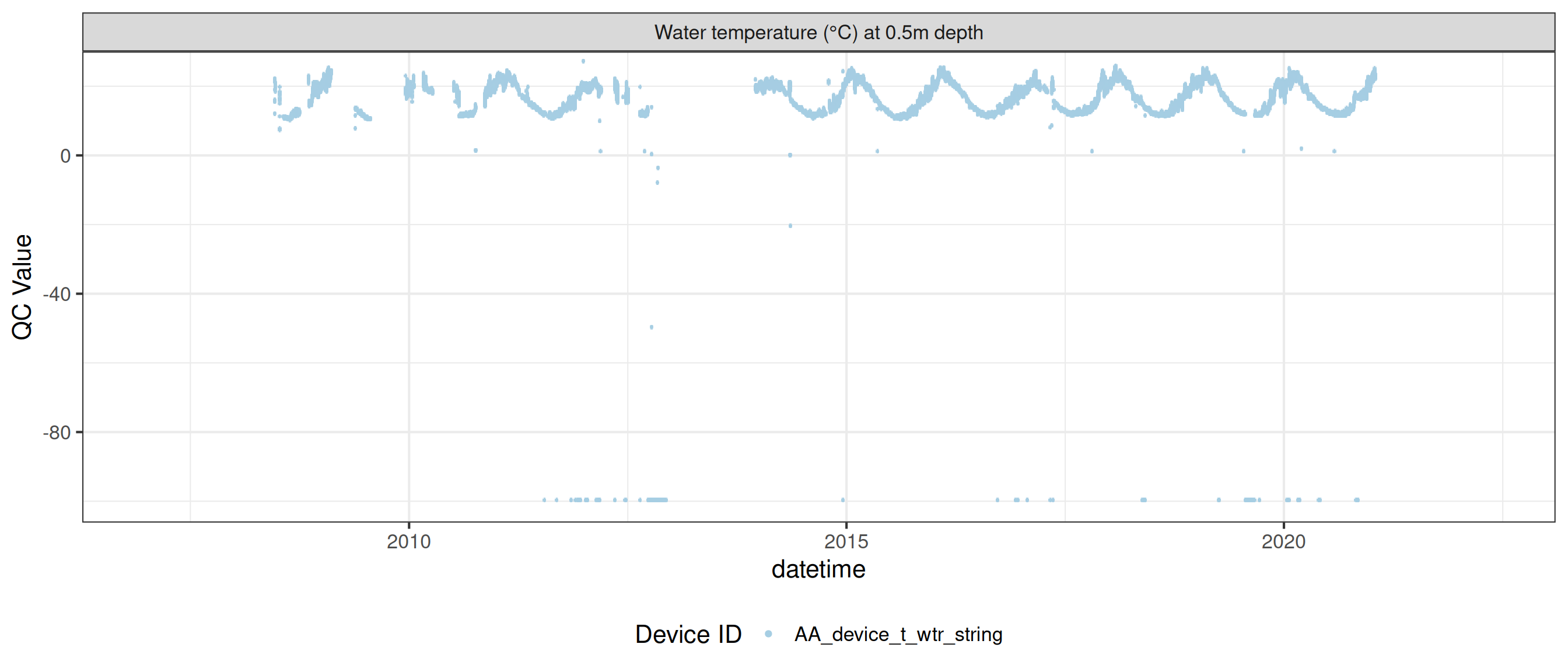
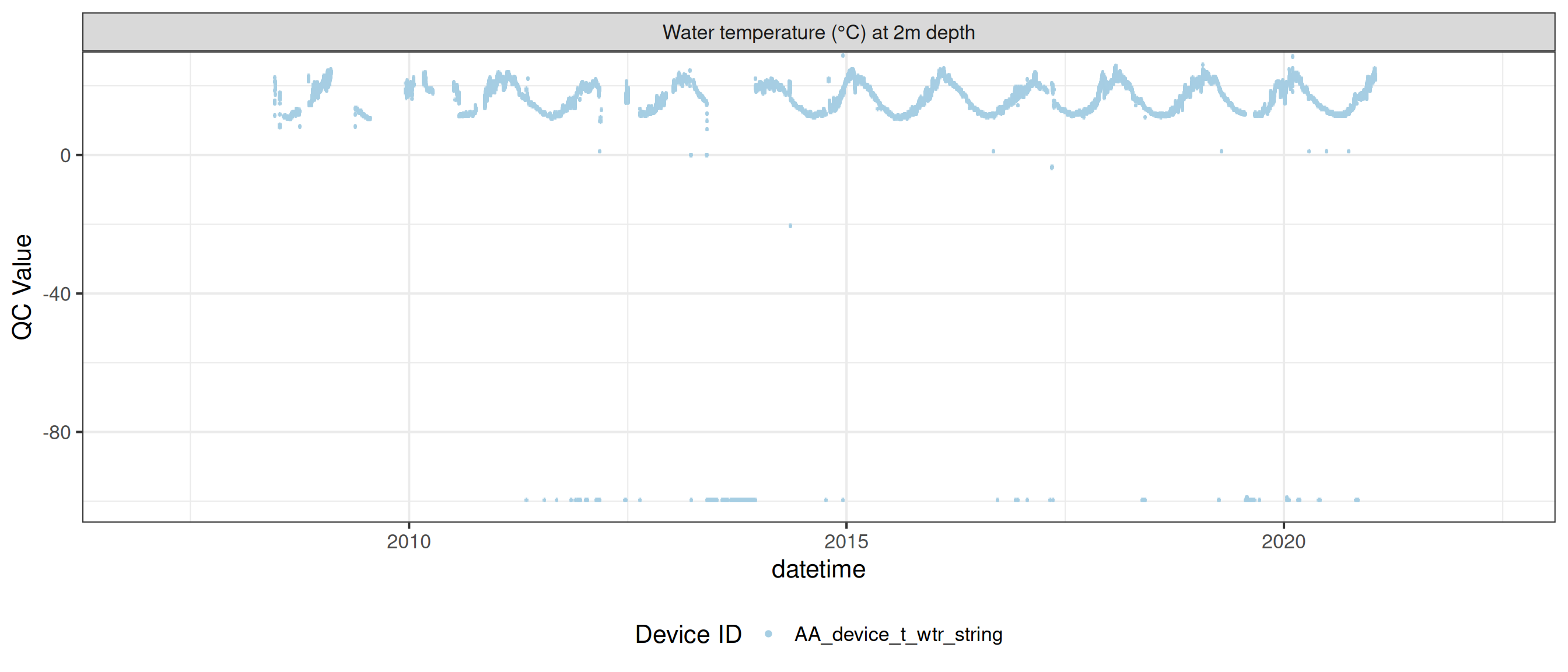
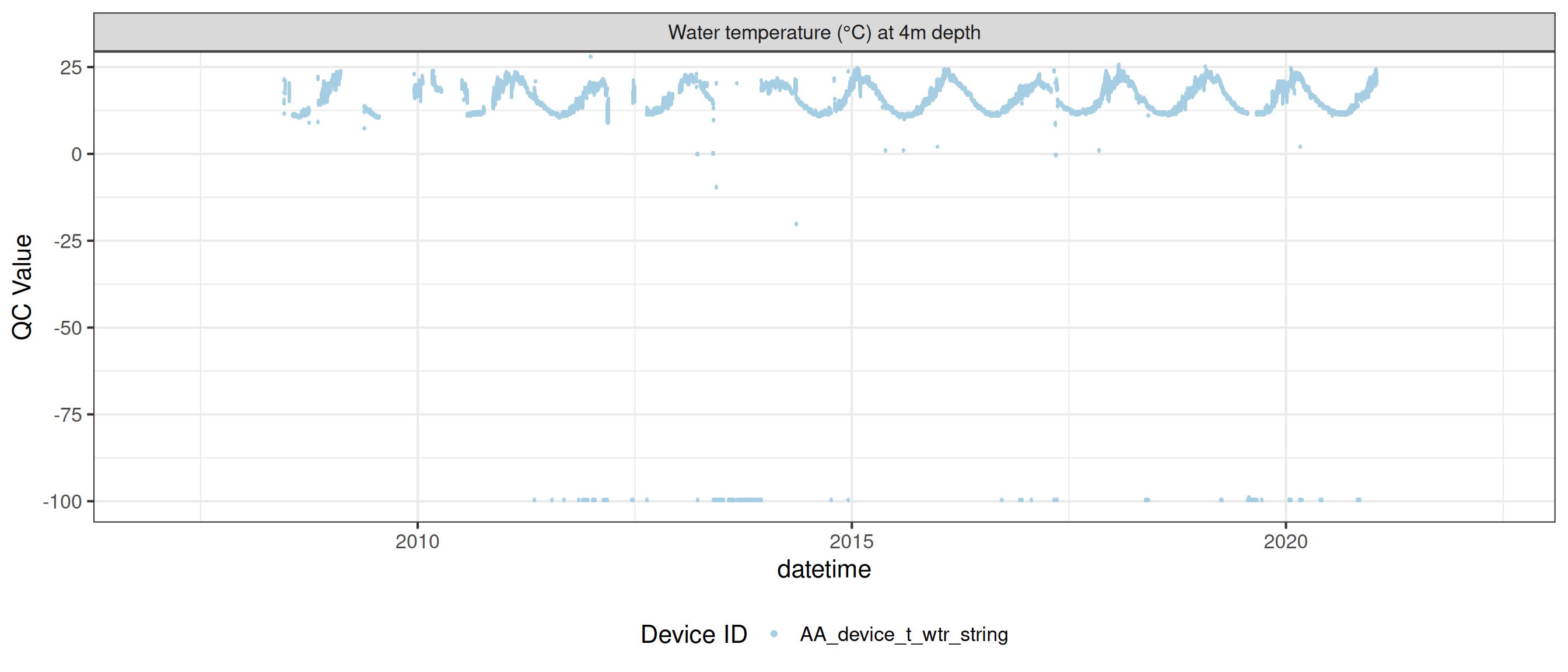
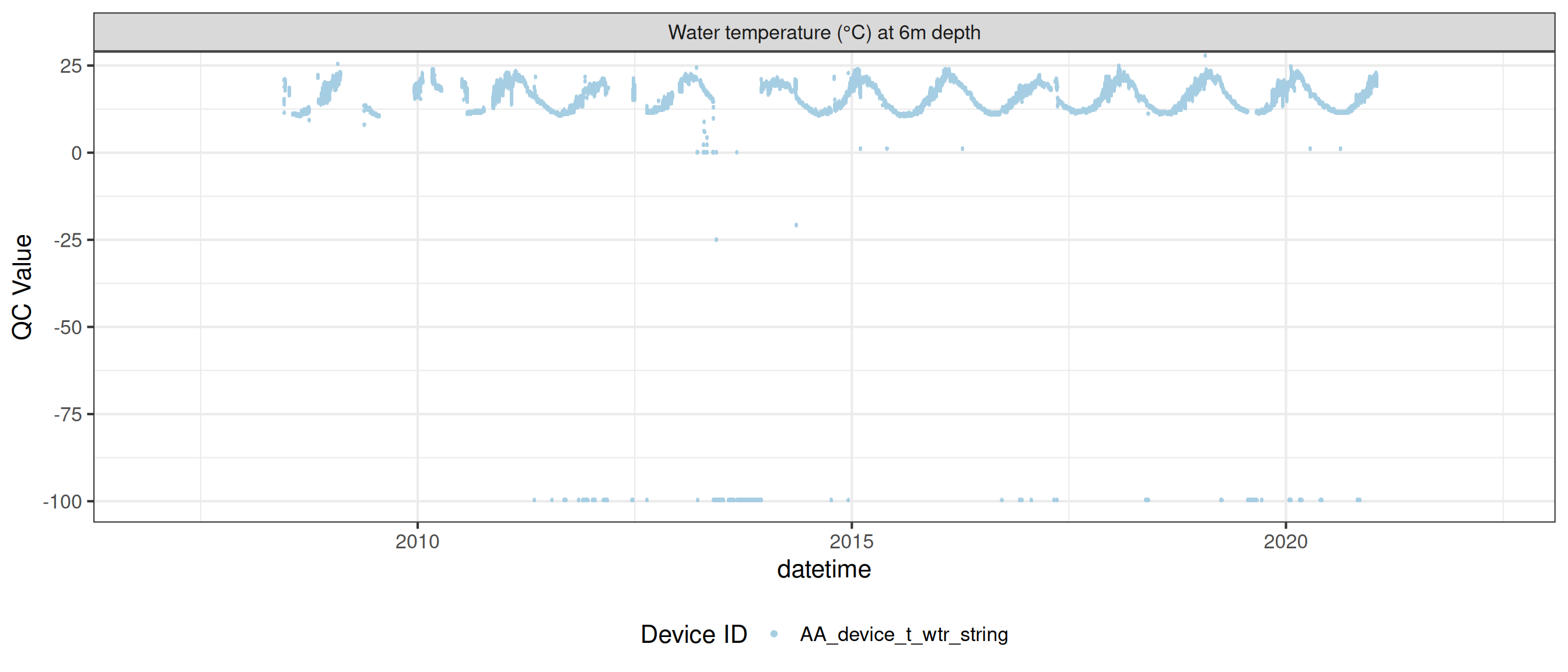
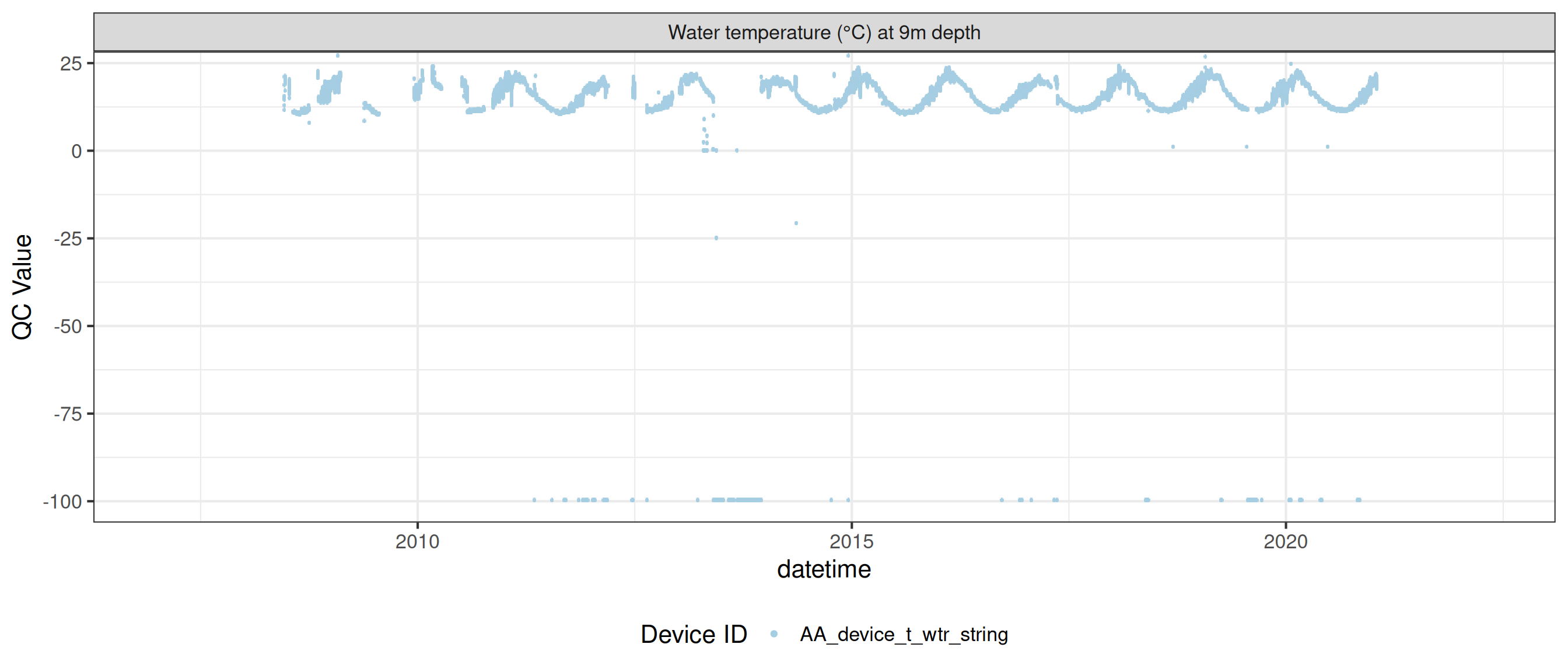
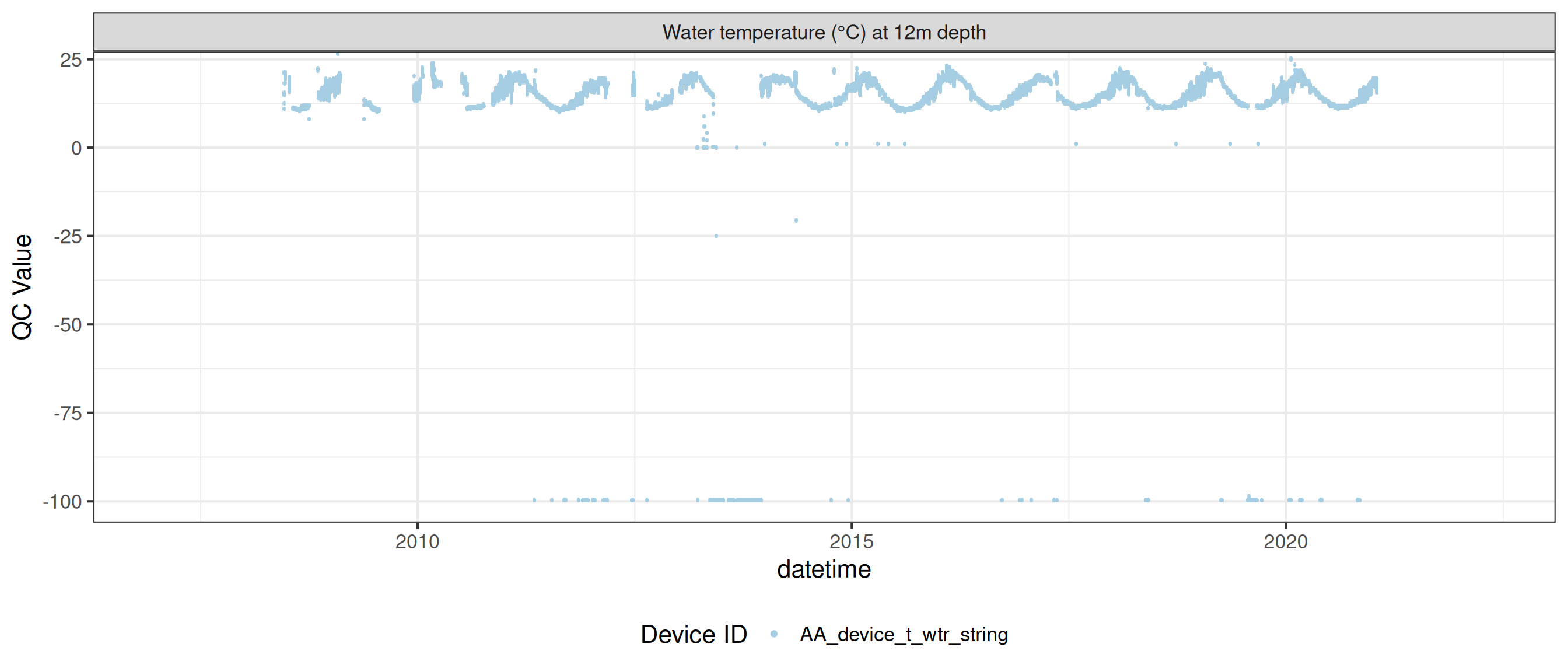
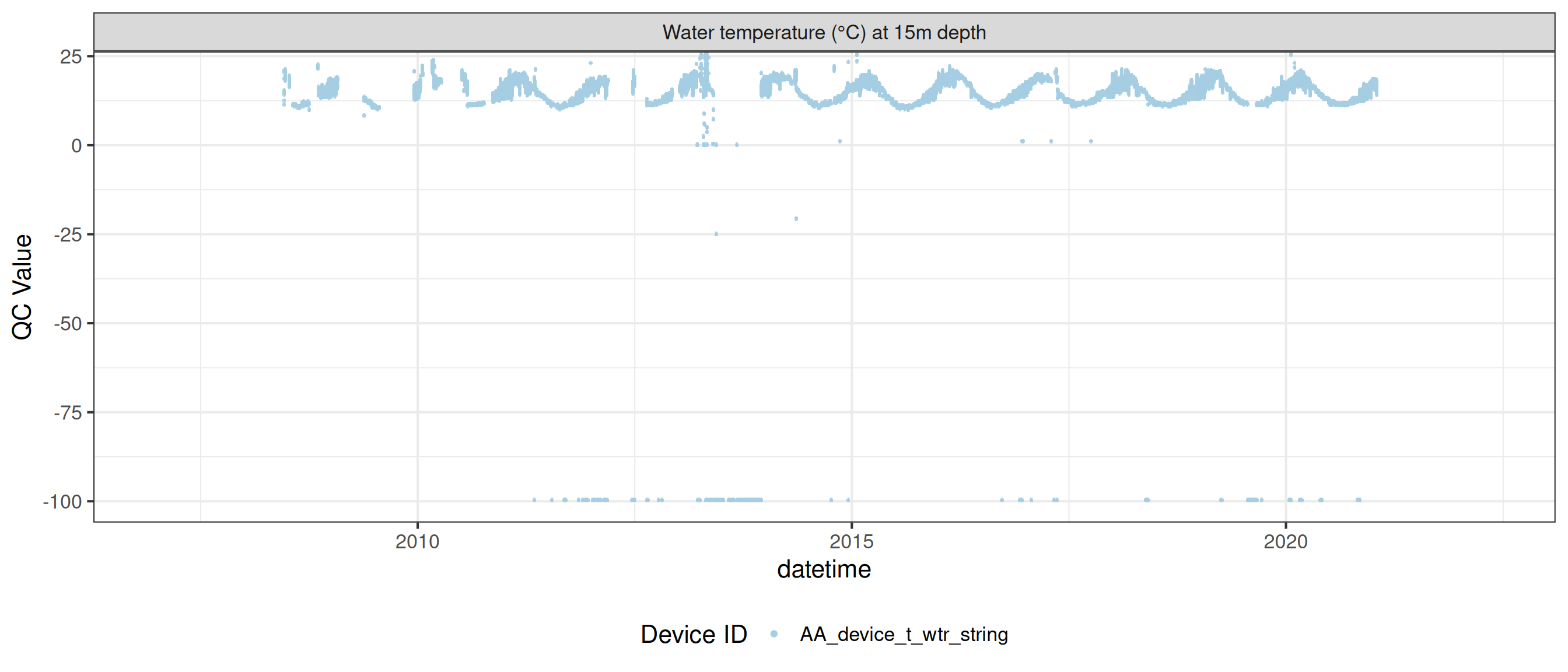
<- raw_long_device_filt |> filter (var_abbr %in% c ("t_wtr" )) |> update_qc_code_vectorized (qc_update_df = qc_update_df)
plot_raw_qc (data = wtemp, variable_ref = variable_ref, ylim = c (7 , 30 ))
Check for offsets between temperature nodes
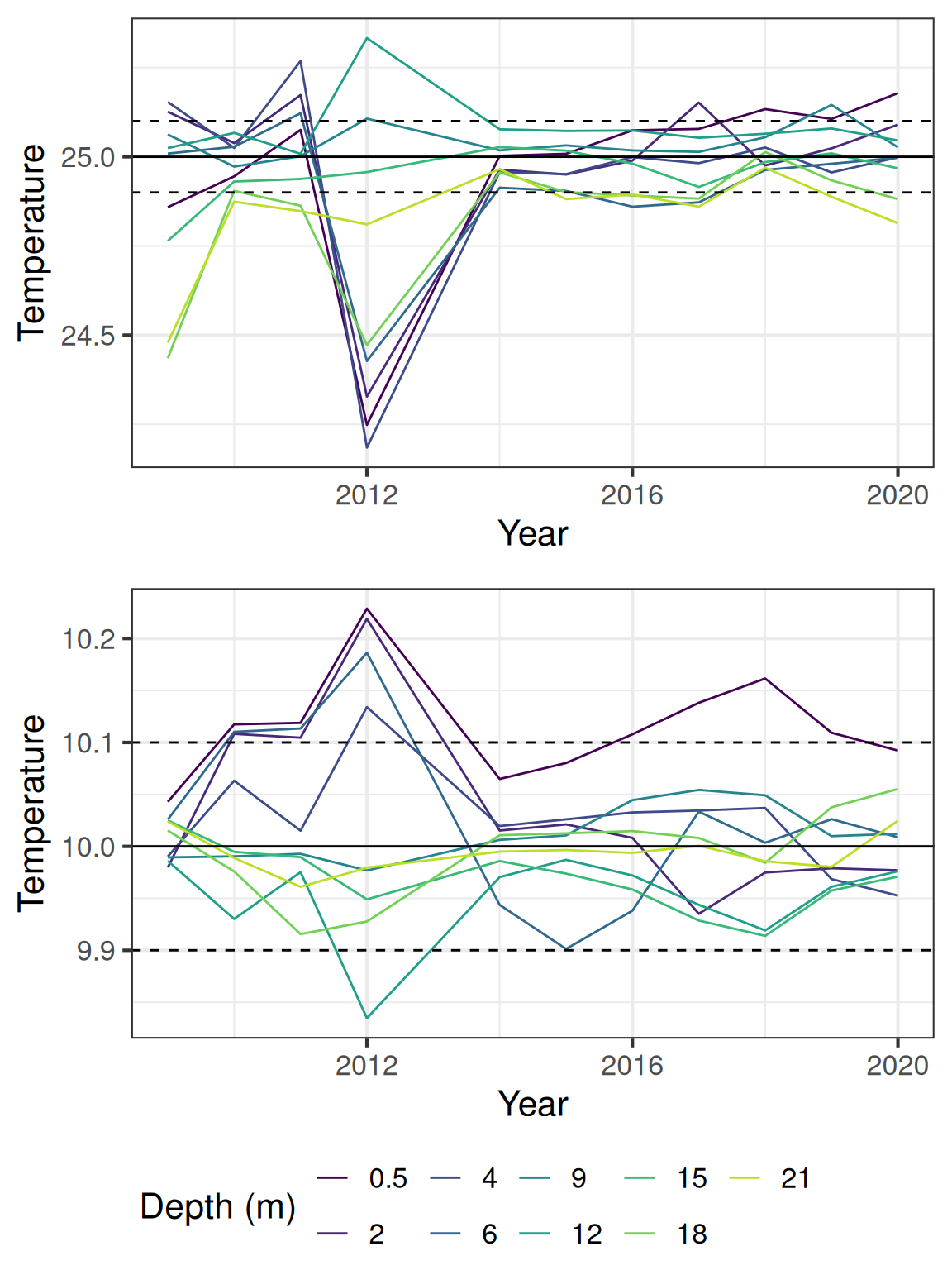
Generally the temperature data falls within the expected range, but there are some offsets between the sensors. The following plot shows the estimated temperatures for 10 and 25 degrees C, using calculated offsets based on when the temperature difference between the sensors are in 5% of the distribution.
The temperature from the DO sensors (1m and 10m) have a lower accuracy (0.5 C).
<- wtemp |> # Filter out oxygen depths filter (! var_ref_id %in% c ("t_wtr_d1810" )) |> pull (device_id) |> unique ()<- site_devices |> filter (device_id %in% temp_strings)
Assign QC codes for met variables
<- raw_long_device_filt |> filter (var_abbr %in% c ("w_dir" , "w_spd" ))
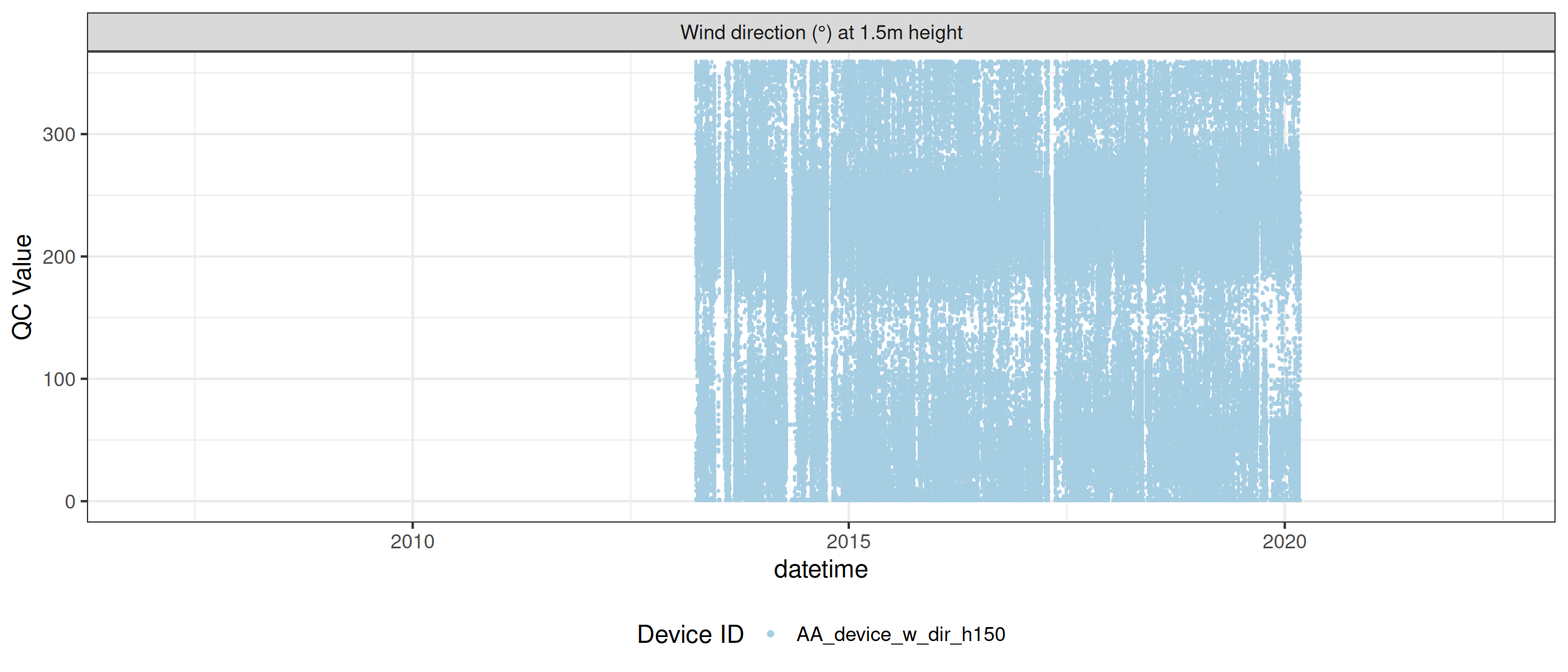
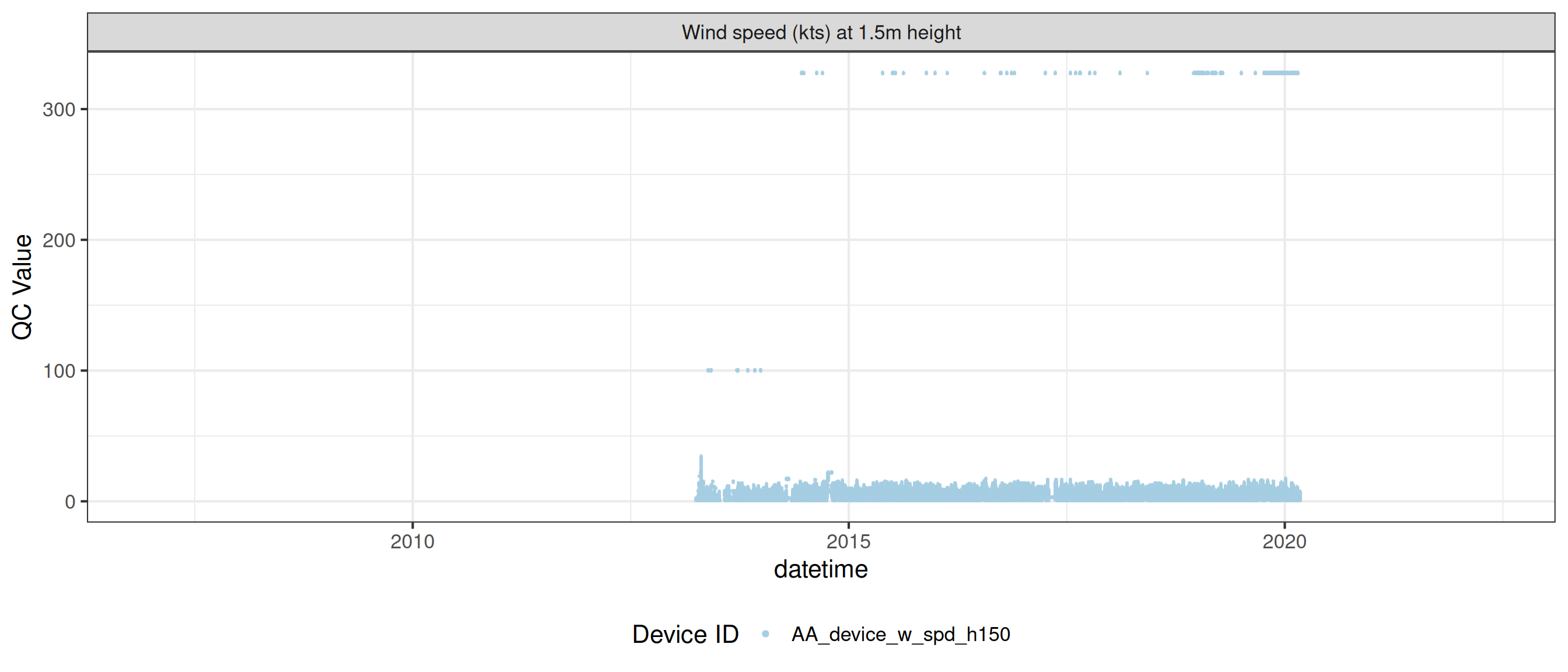
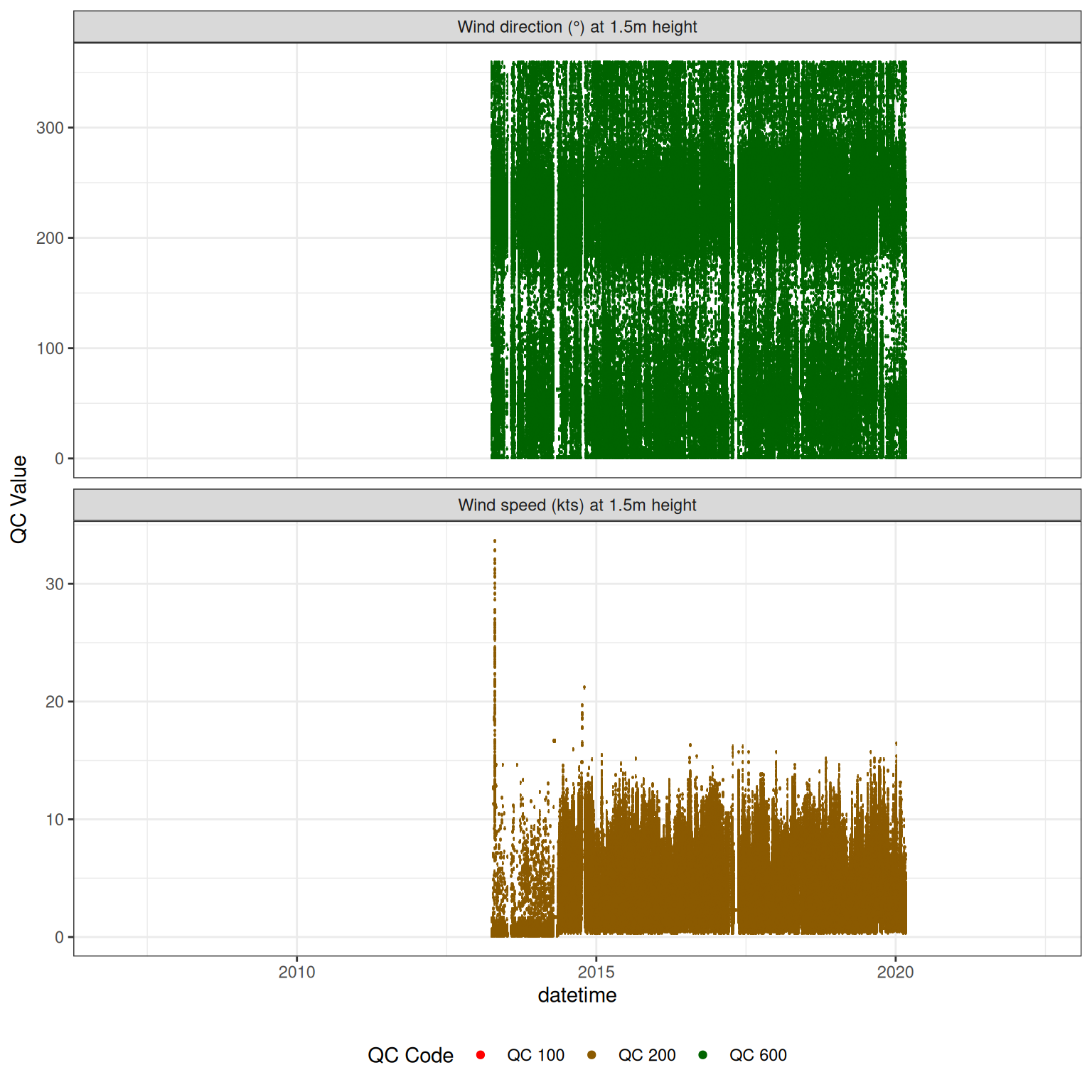
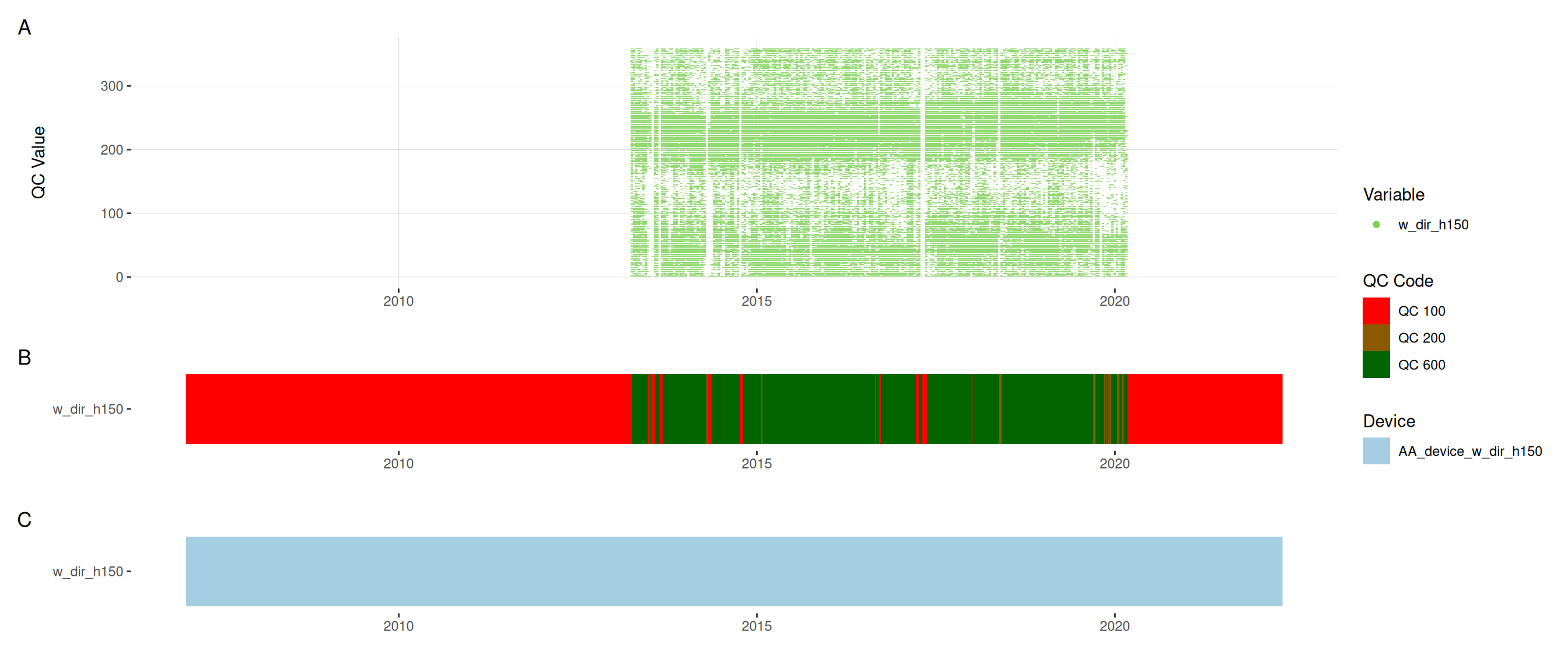
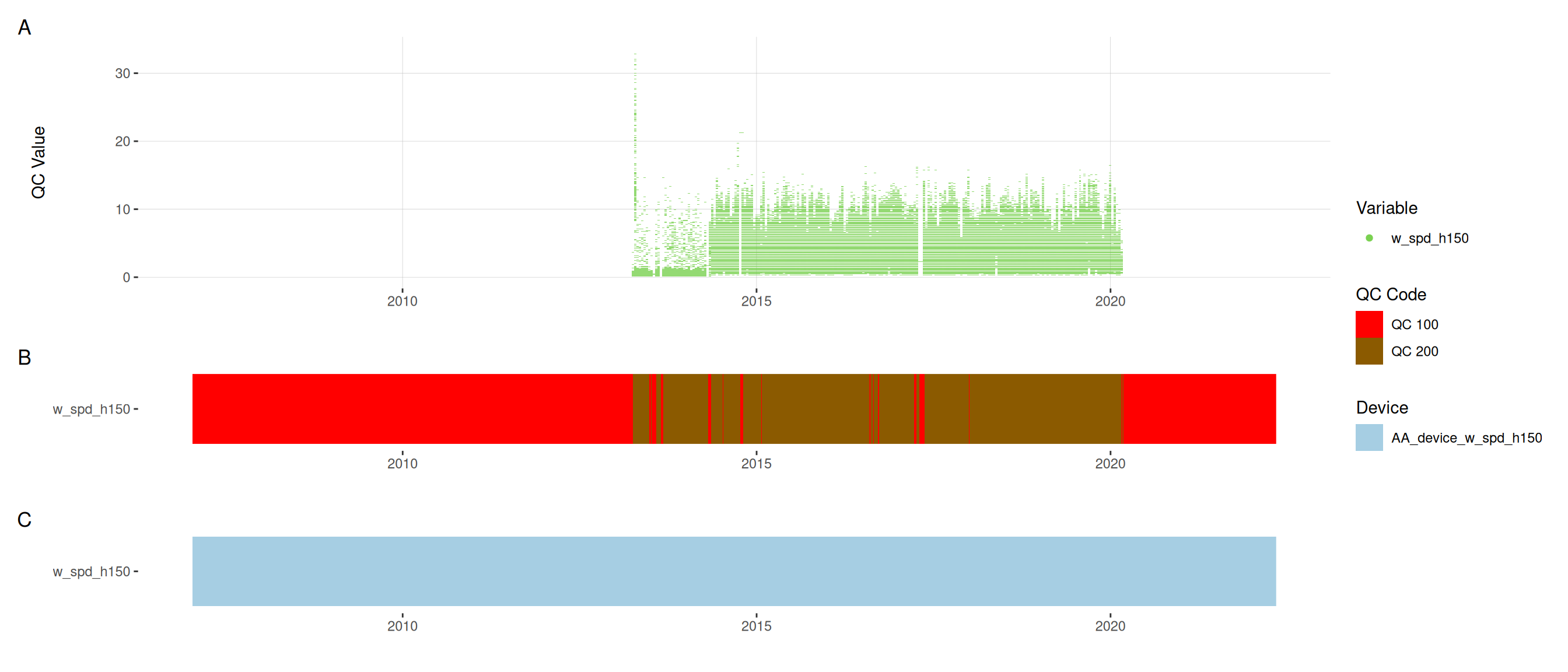
<- met |> update_qc_code_vectorized (qc_update_df = qc_update_df, var_ref_id = c ("w_dir_h150" , "w_spd_h150" ))|> plot_sensor (var_ref_id = c ("w_dir_h150" , "w_spd_h150" ), variable_ref = variable_ref,sensor_calibrations = sensor_calibrations, sensor_reference = sensor_reference,sensor_scaling = sensor_scaling, clip_ylim = FALSE , colour = "qc_code" )
plot_raw_qc (data = met, variable_ref = variable_ref)
Visual summaries
Plot each variable
Temperature
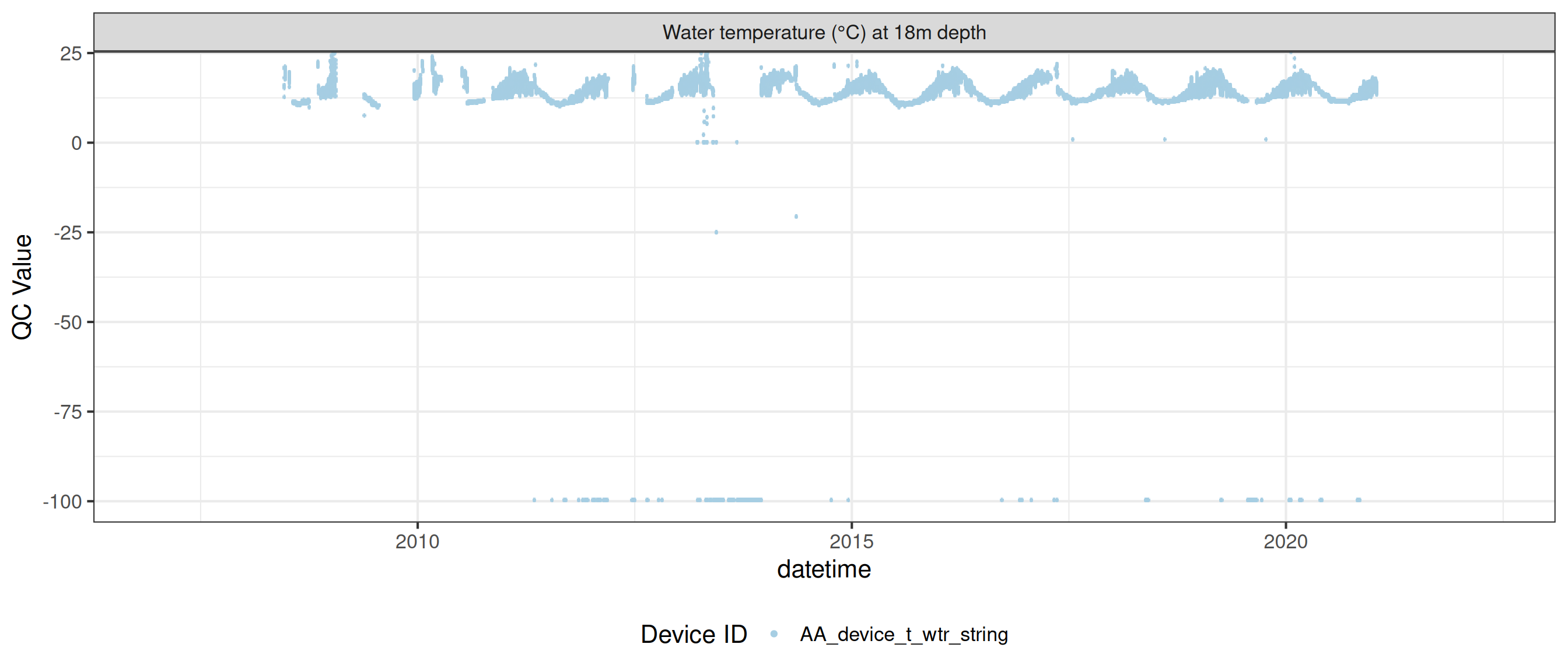
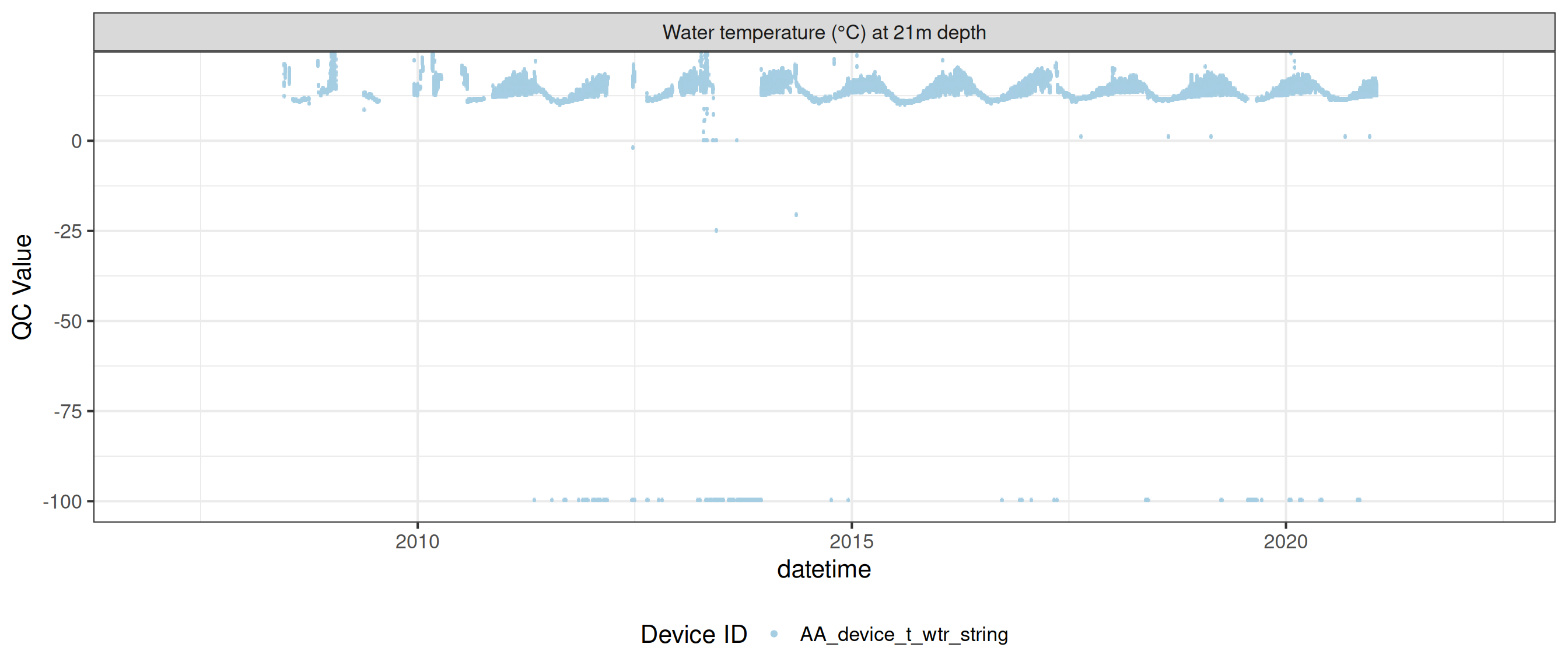
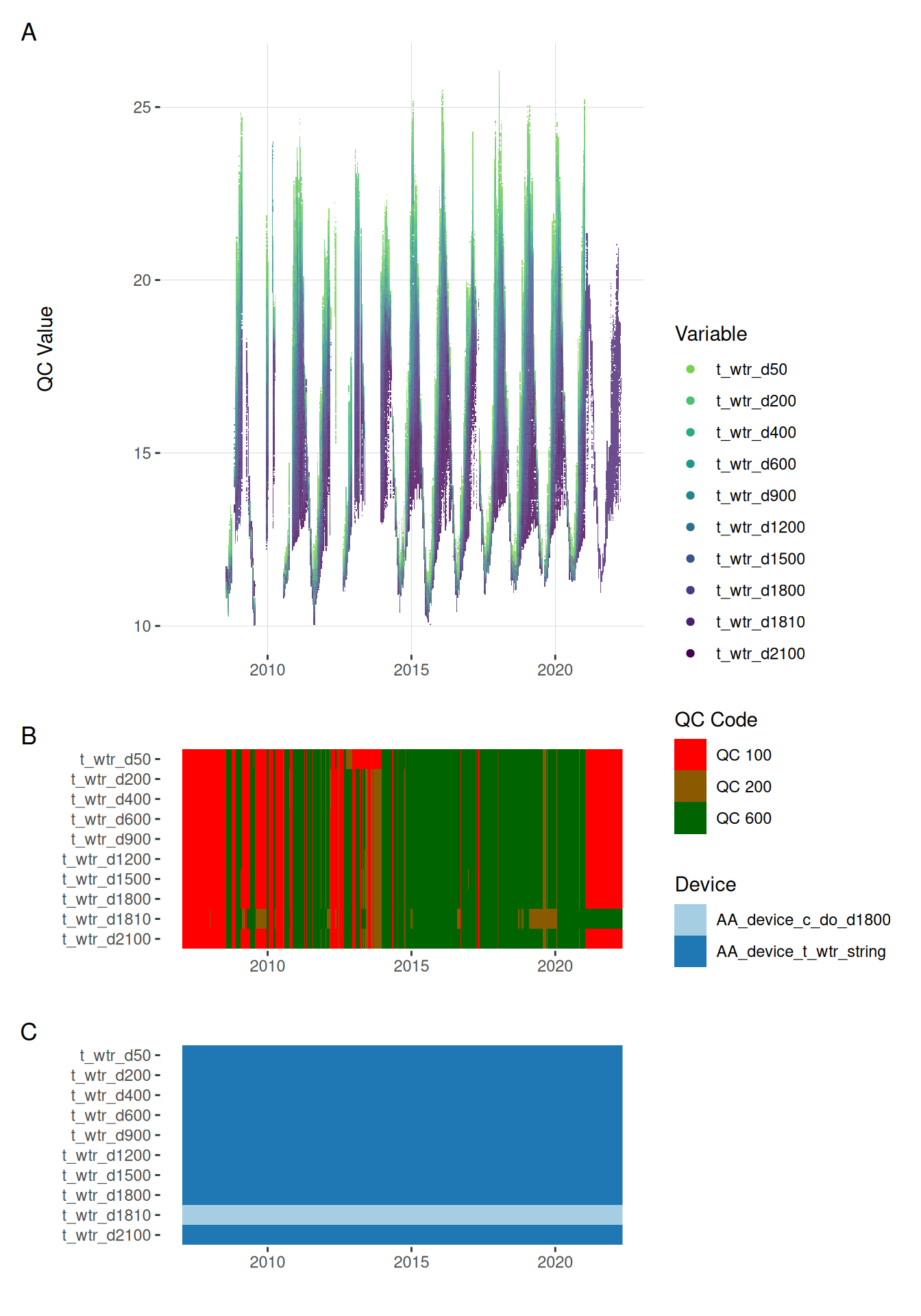
plot_var_ts_qc (data = data, var_ref_id = c ("t_wtr_d50" , "t_wtr_d200" ,"t_wtr_d400" , "t_wtr_d600" ,"t_wtr_d900" , "t_wtr_d1200" ,"t_wtr_d1500" , "t_wtr_d1800" , "t_wtr_d1810" , "t_wtr_d2100" ))
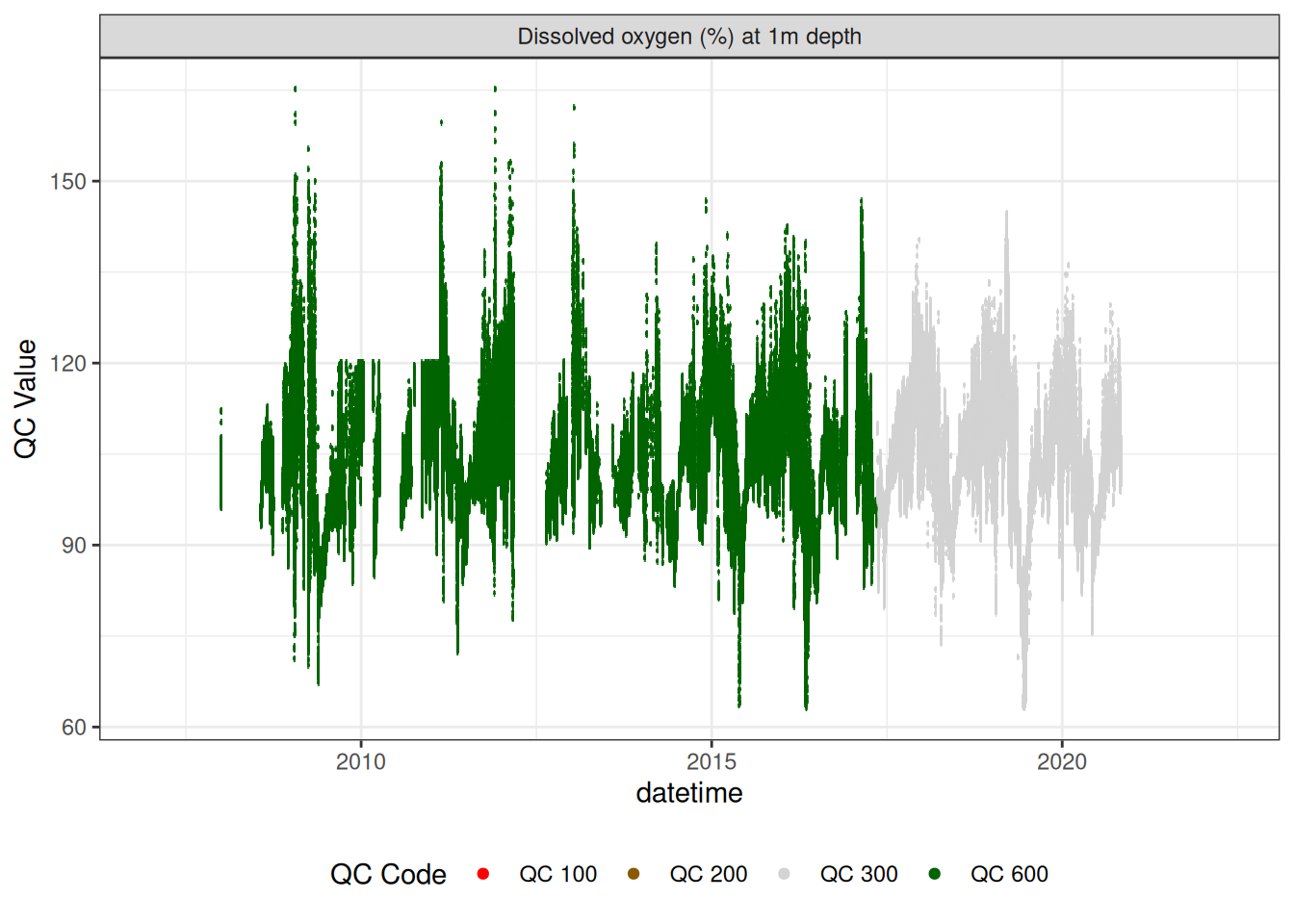
Oxygen
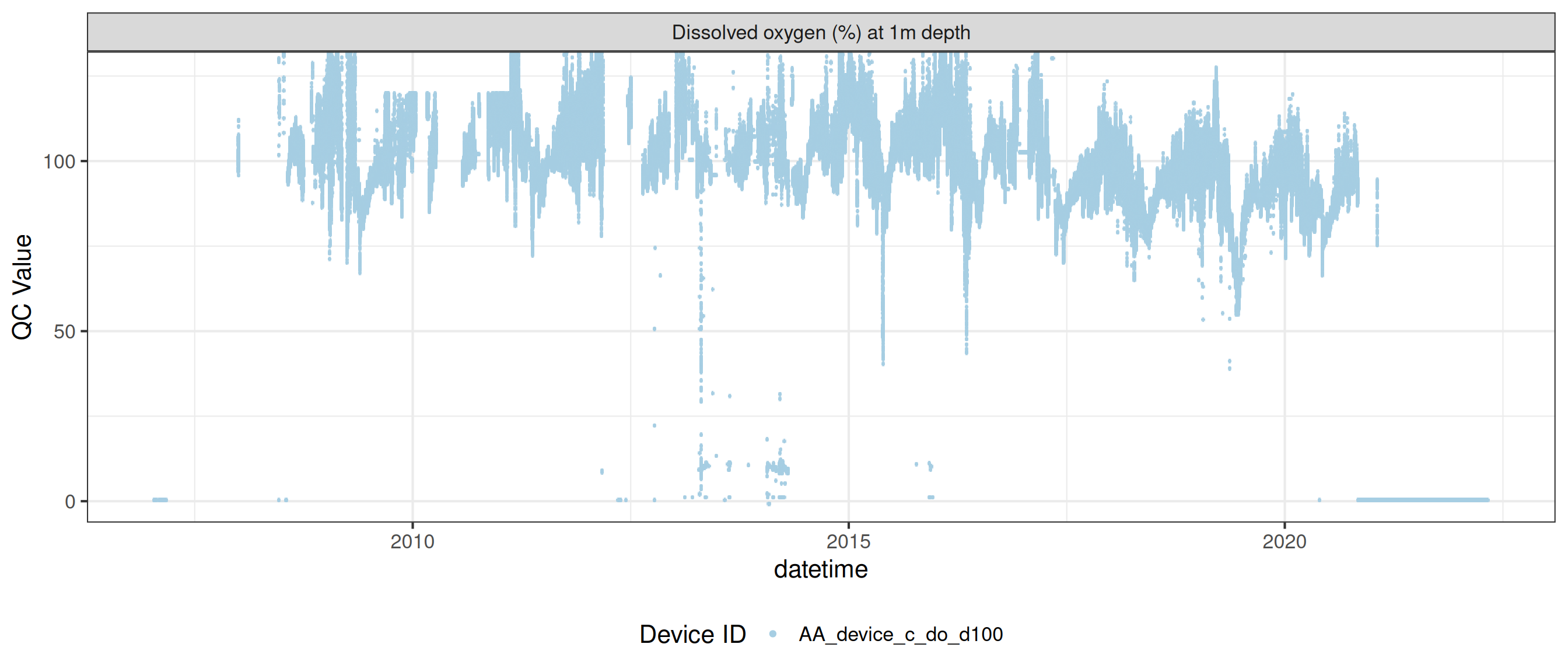
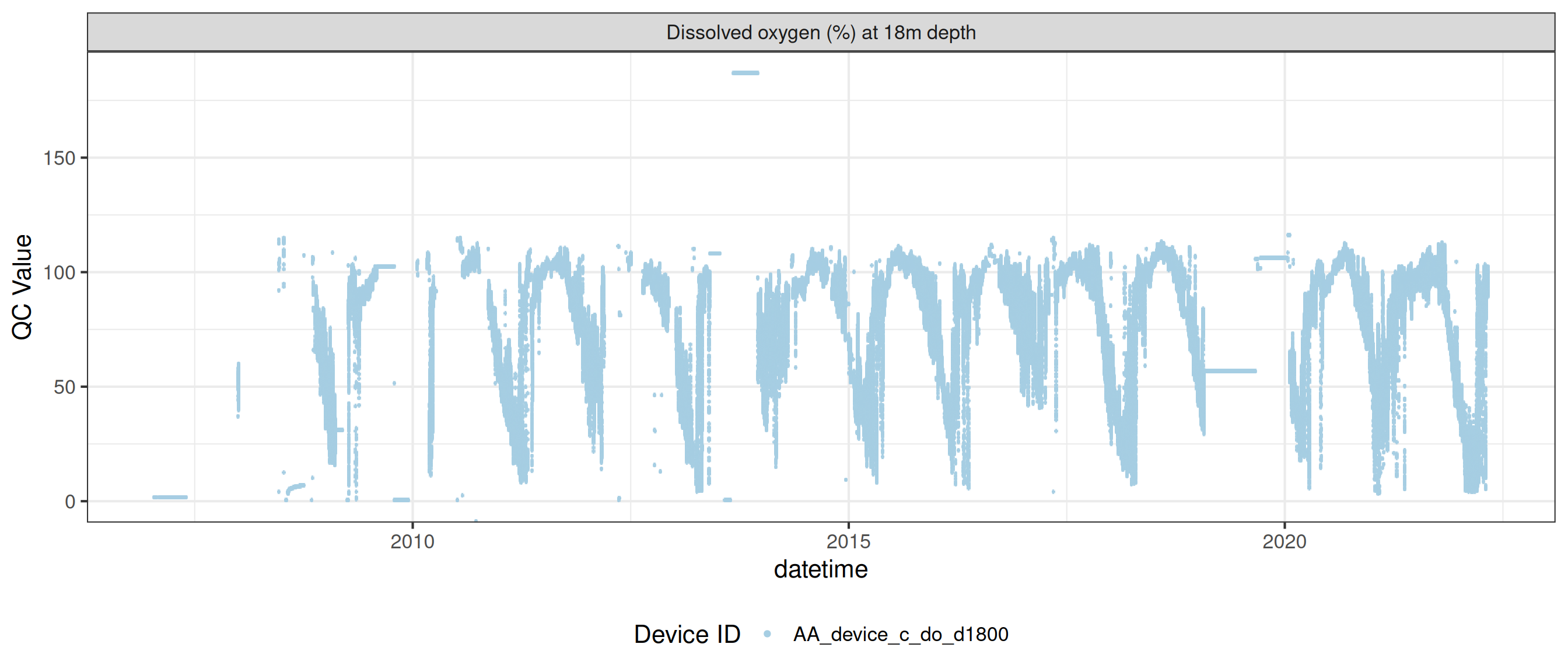
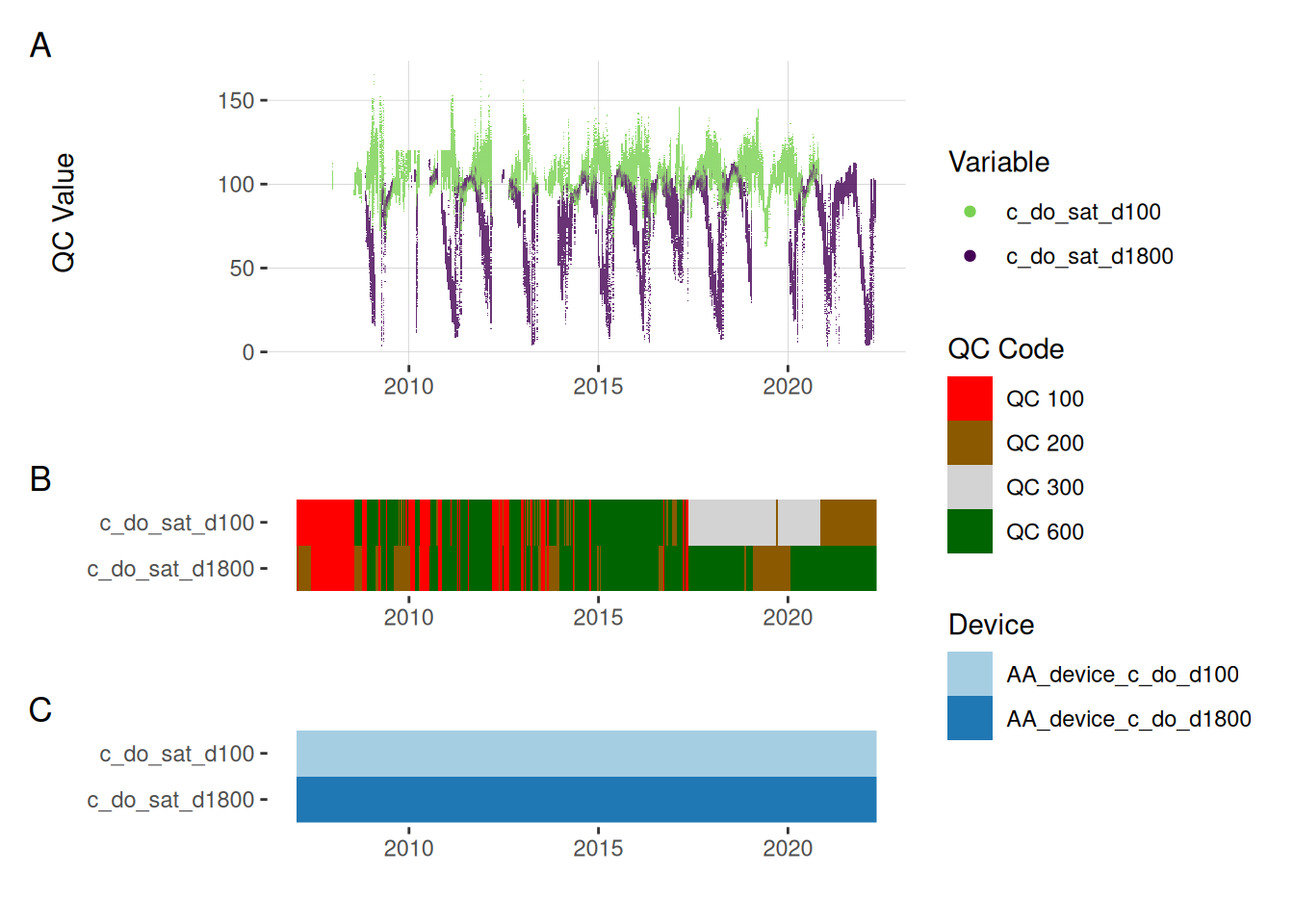
plot_var_ts_qc (data = data, var_ref_id = c ("c_do_sat_d100" , "c_do_sat_d1800" ))
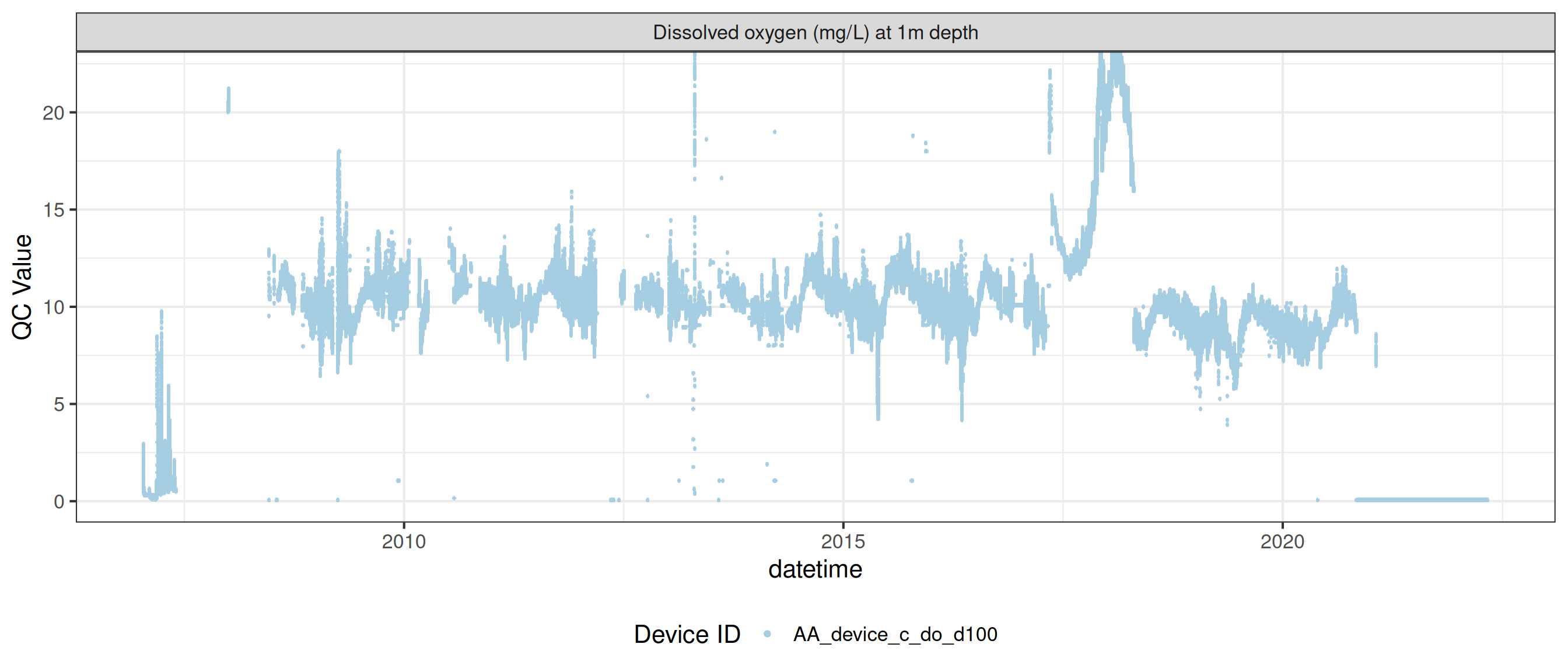
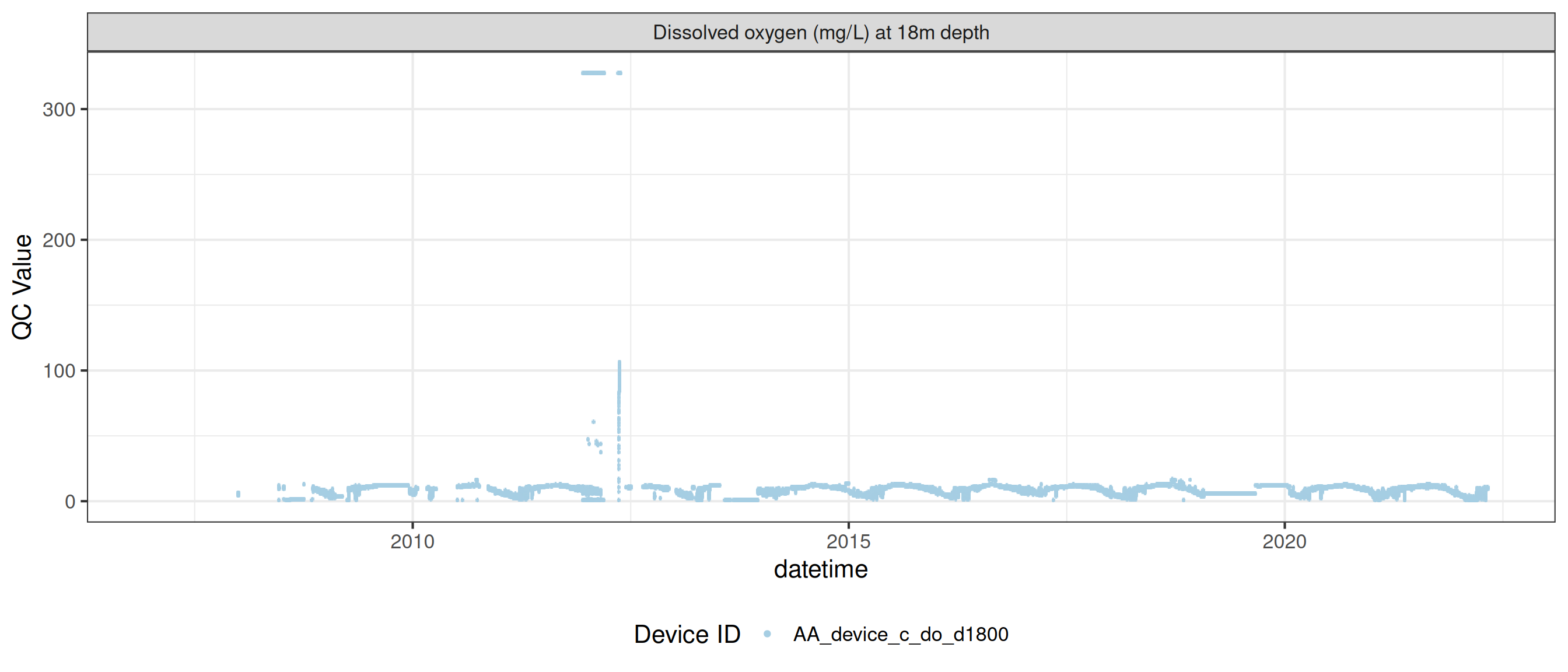
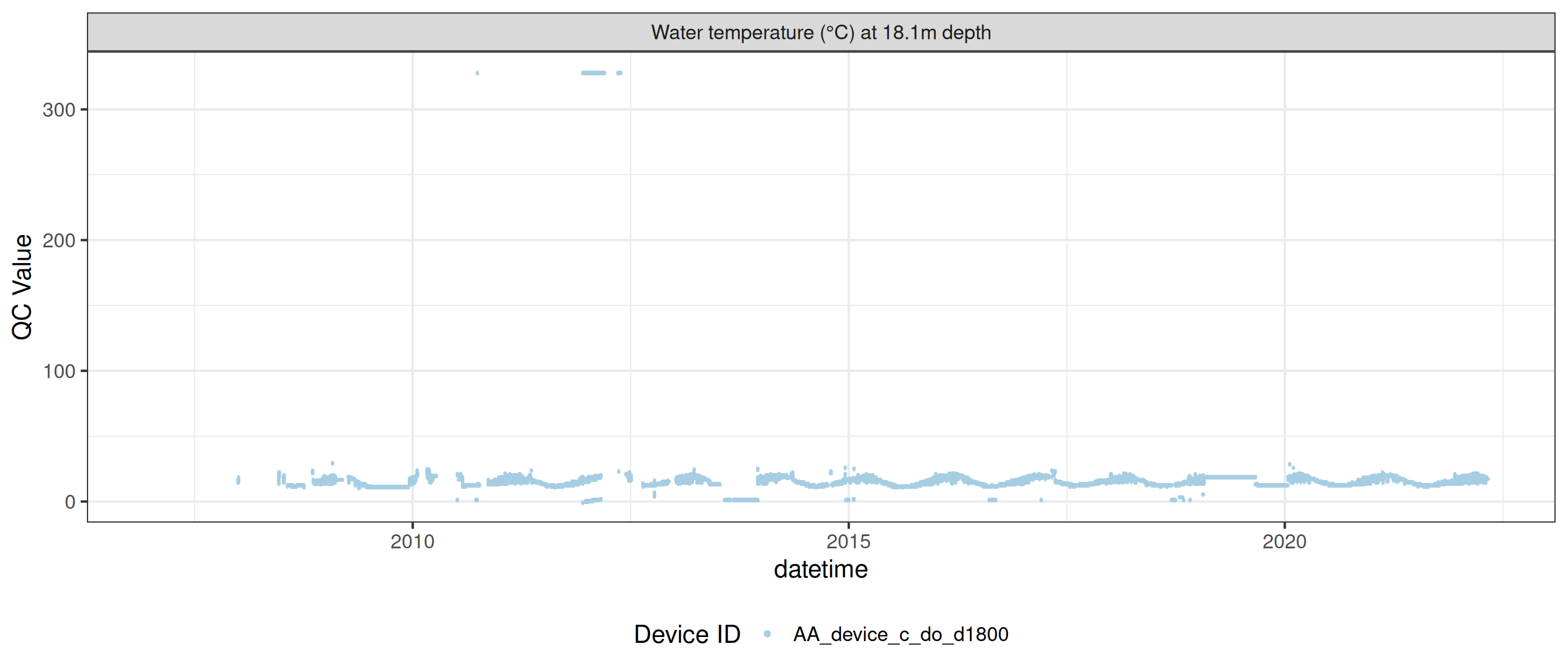
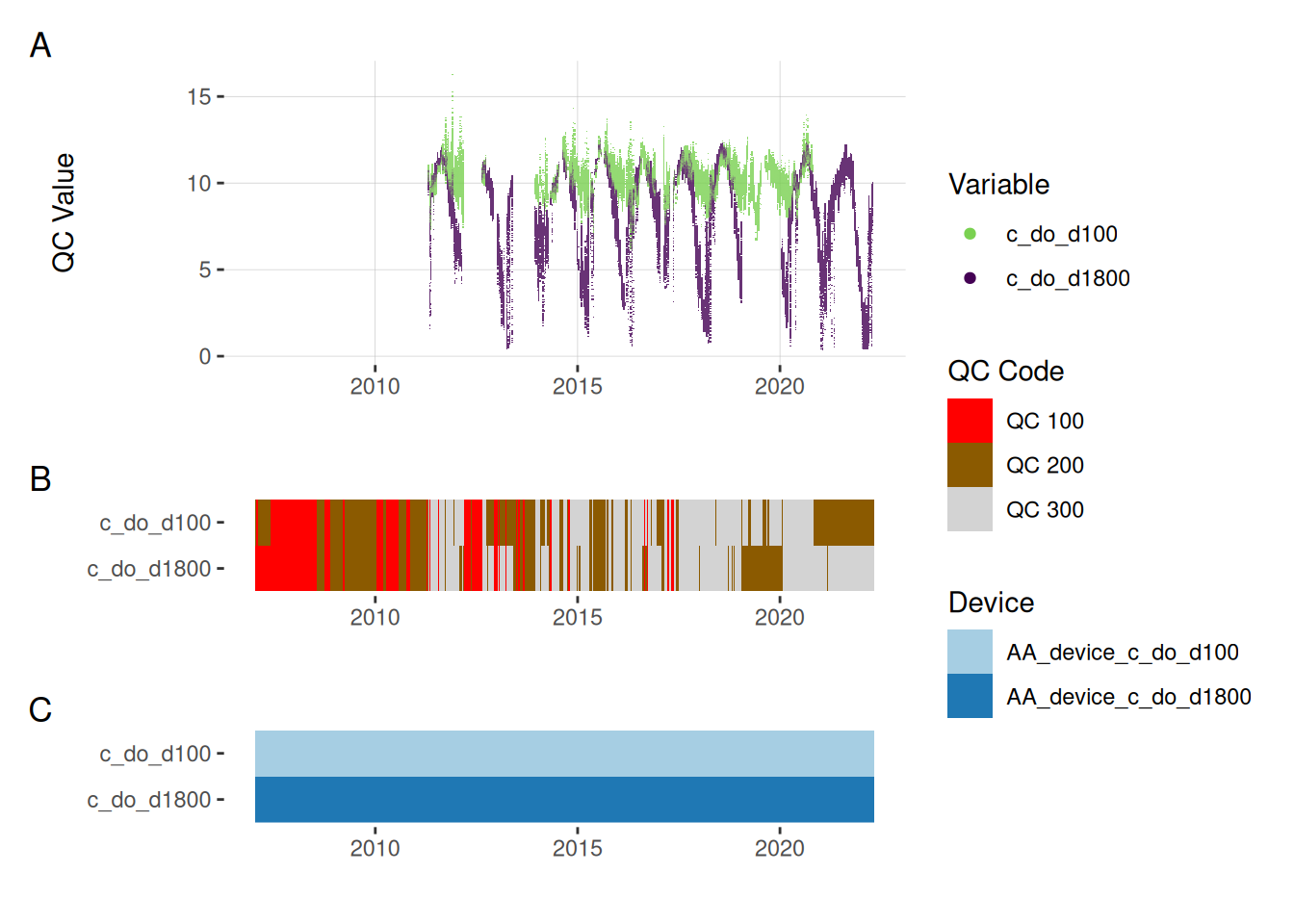
plot_var_ts_qc (data = data, var_ref_id = c ("c_do_d100" , "c_do_d1800" ))
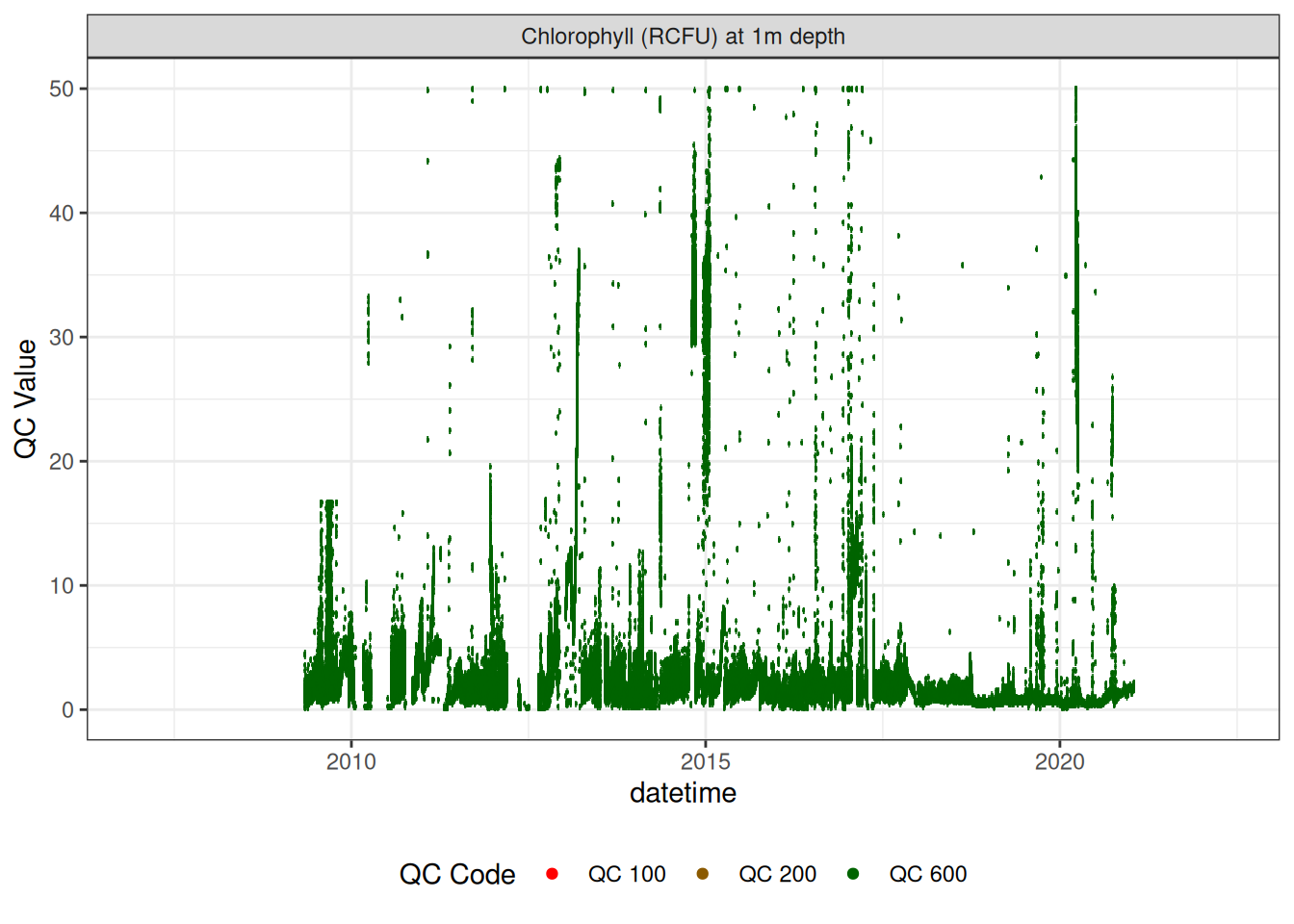
Chlorophyll
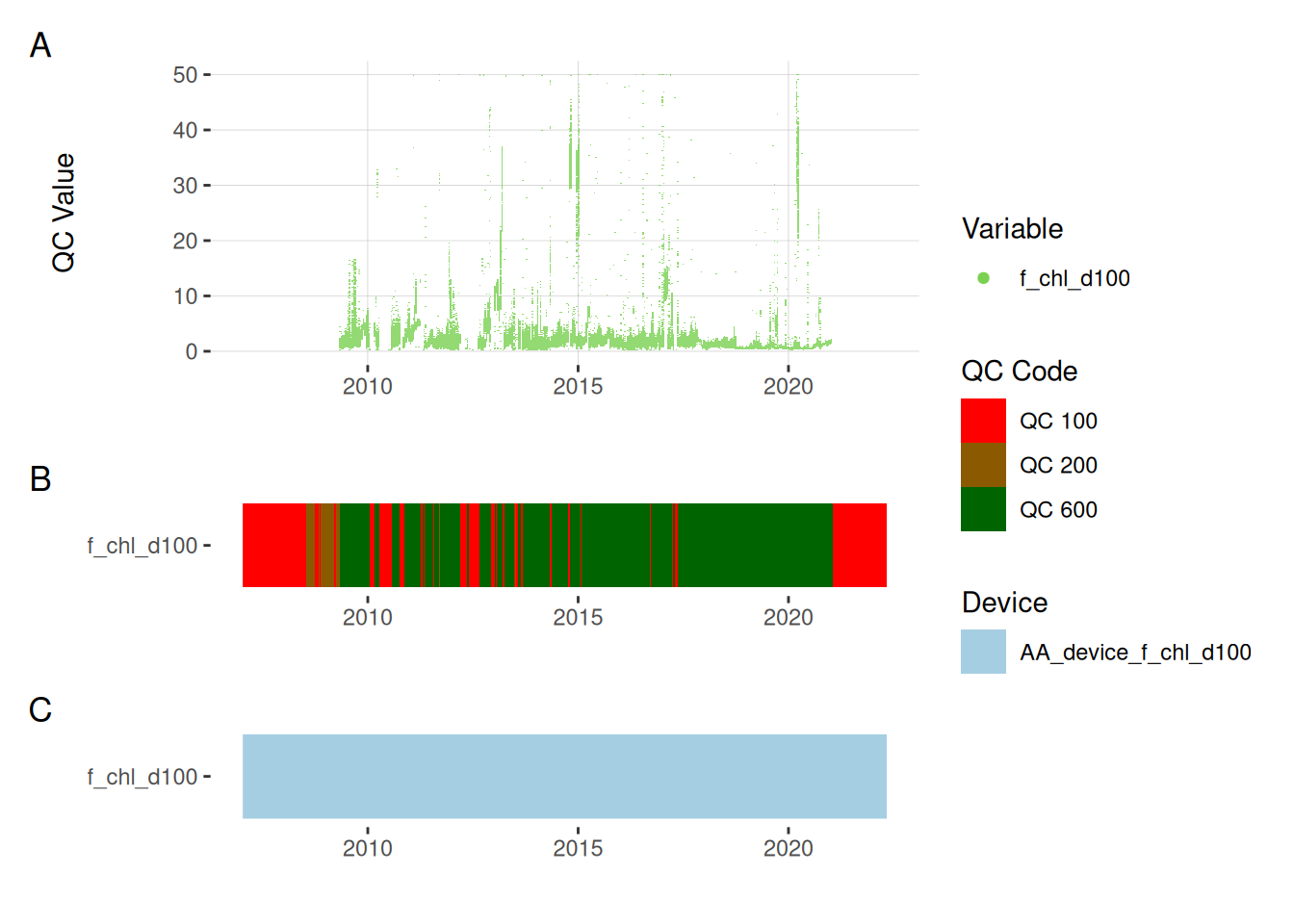
plot_var_ts_qc (data = data, var_ref_id = c ("f_chl_d100" ))
Download data
Download the data as a zip folder. The data is in wide format an comes with metadata files. The metadata files contain the information needed to reconstruct the quality control.
Download Data